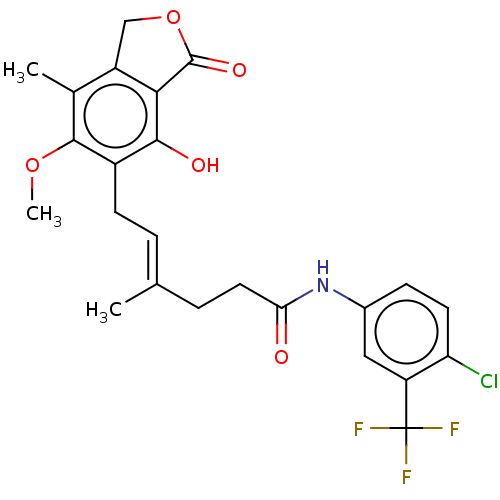
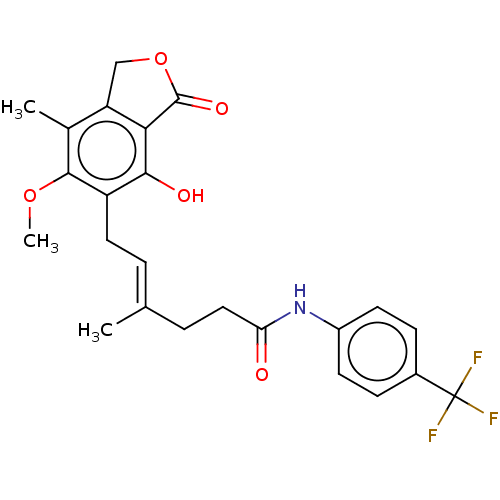
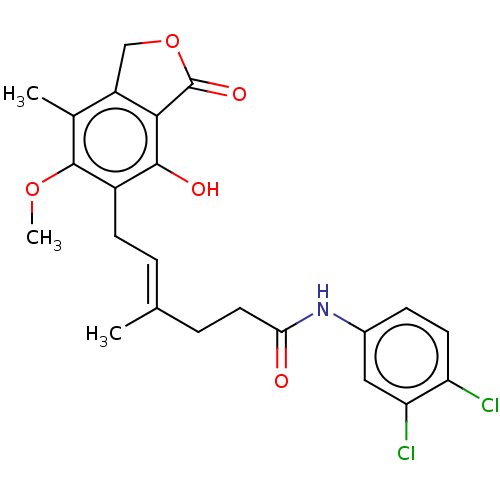
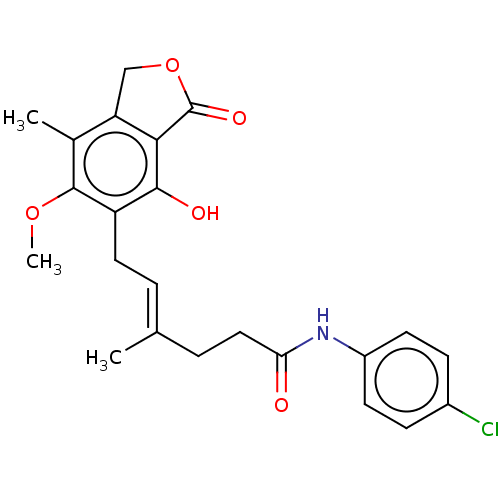
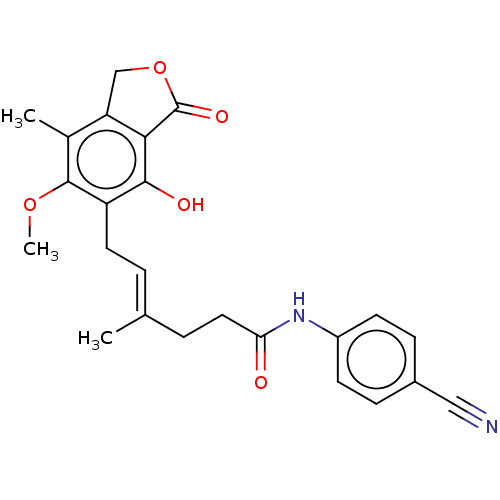
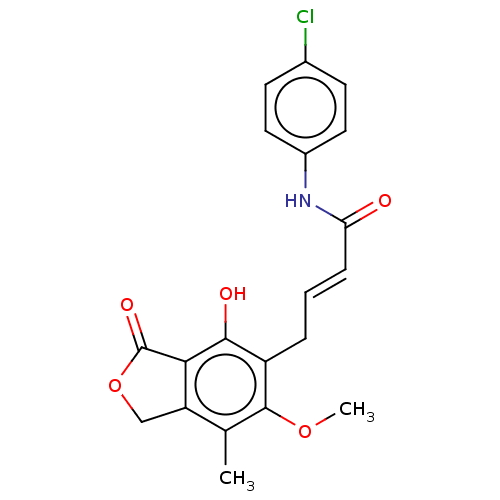
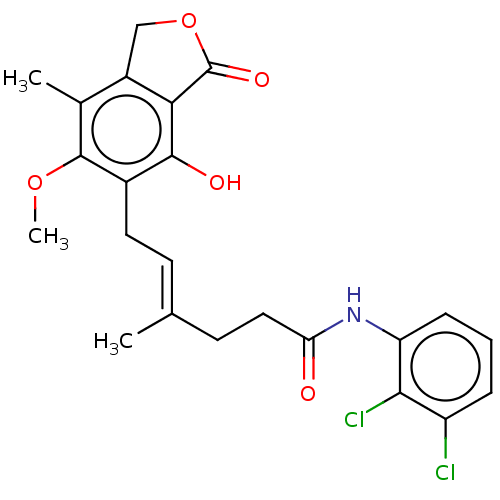
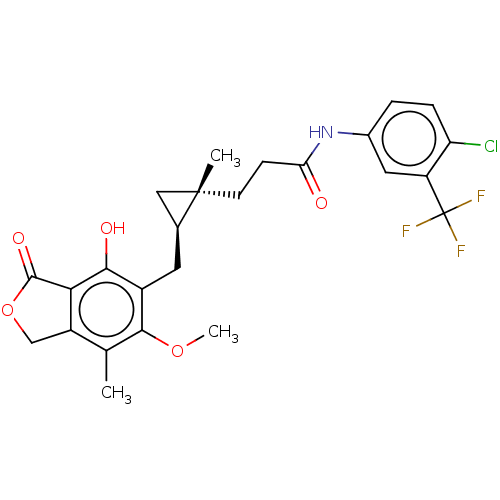
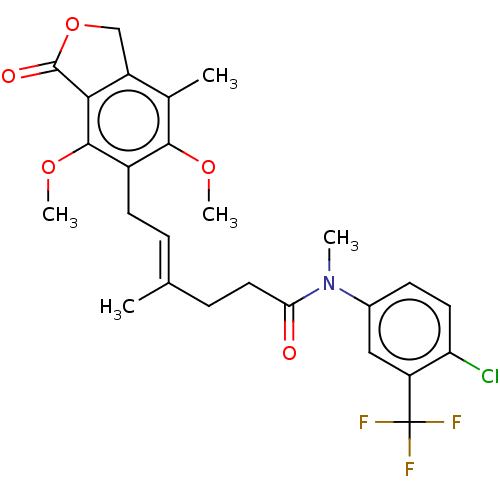
Report error Found 30 Enz. Inhib. hit(s) with all data for entry = 50011548
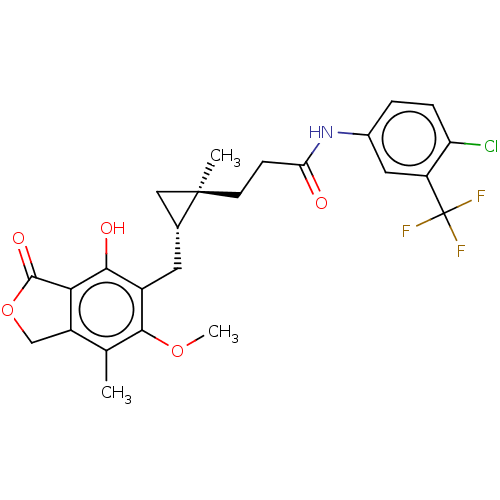
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 16nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
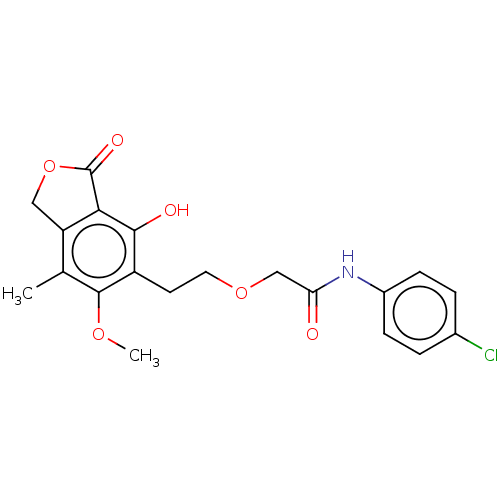
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 40nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
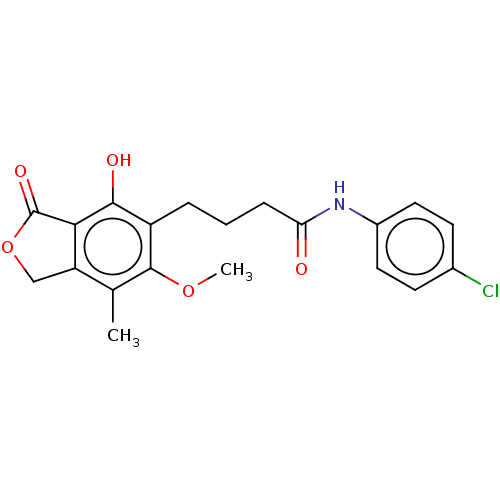
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
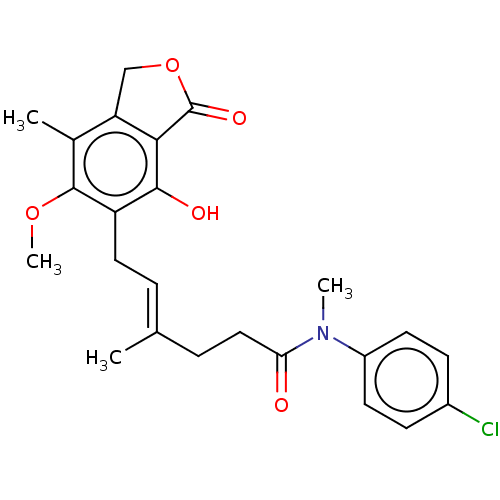
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 42nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 46nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 60nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 60nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 66nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 130nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 143nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 176nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
Affinity DataKi: 230nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 270nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 350nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 350nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 460nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 480nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
Affinity DataKi: 550nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 570nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 870nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 870nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Inhibition of human IMPDH2 expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluorescence assayMore data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 1.70E+3nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair
TargetInosine-5'-monophosphate dehydrogenase(Cryptosporidium parvum)
University of Houston
Curated by ChEMBL
University of Houston
Curated by ChEMBL
Affinity DataKi: 2.80E+3nMAssay Description:Inhibition of Cryptosporidium parvum IMPDH expressed in Escherichia coli assessed as apparent inhibition constant using NAD as substrate by fluoresce...More data for this Ligand-Target Pair