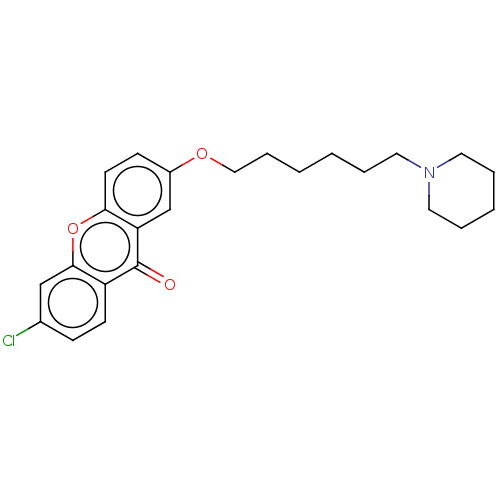
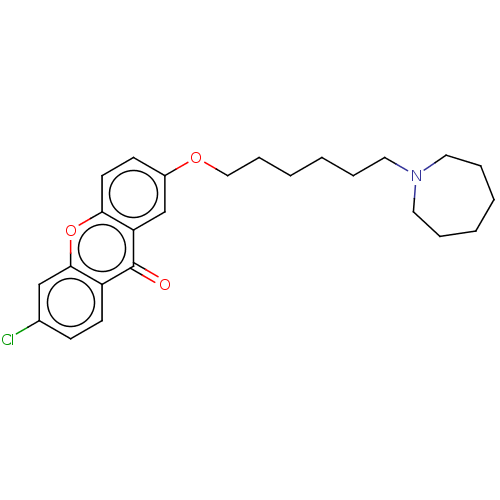
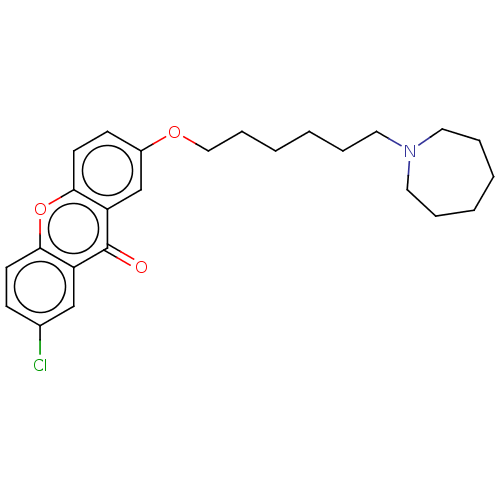
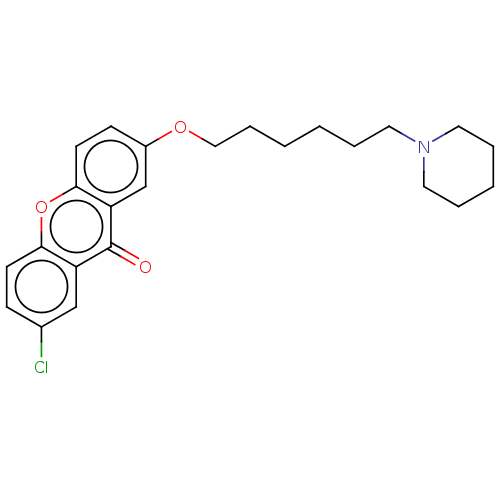
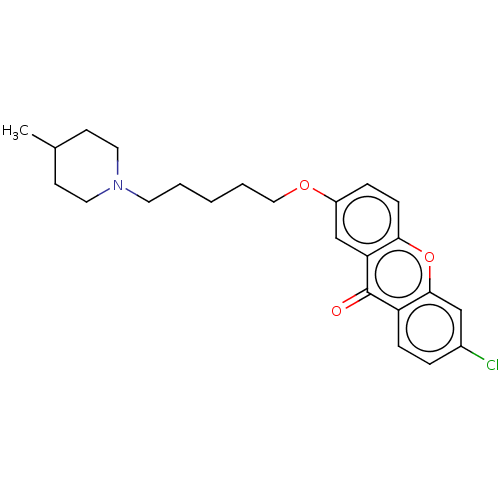
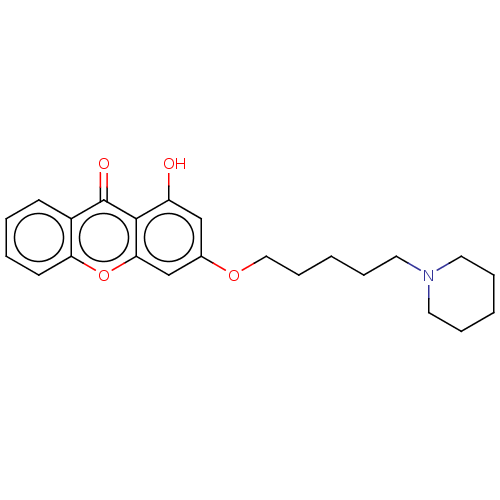
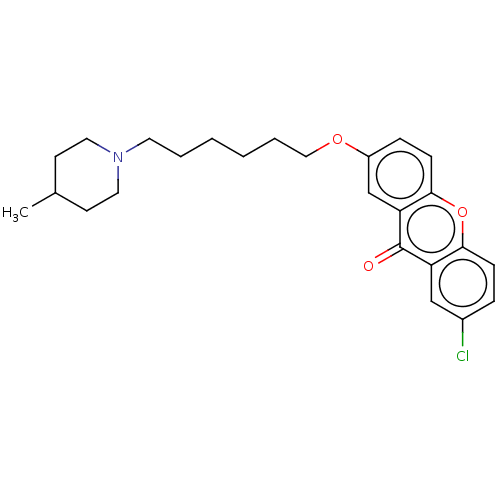
Report error Found 109 Enz. Inhib. hit(s) with all data for entry = 50012274
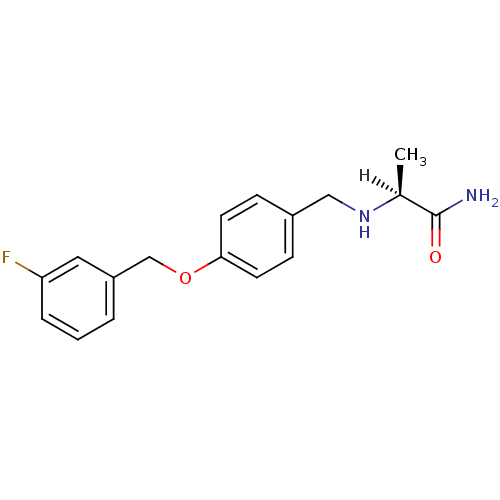
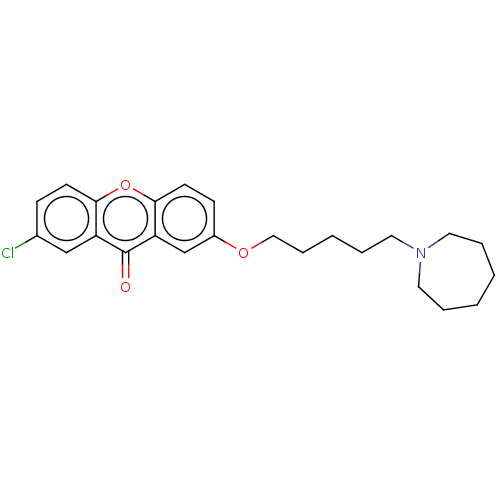
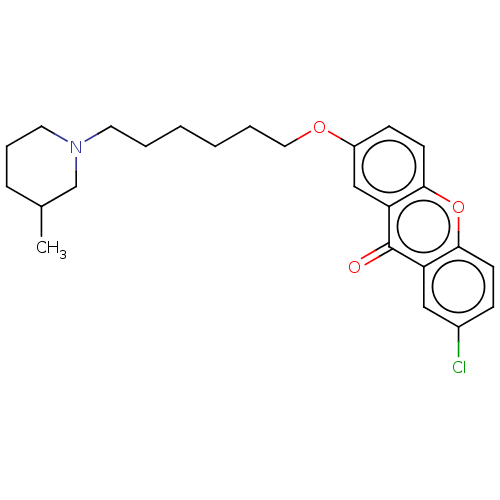
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectro...More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] B(Human)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
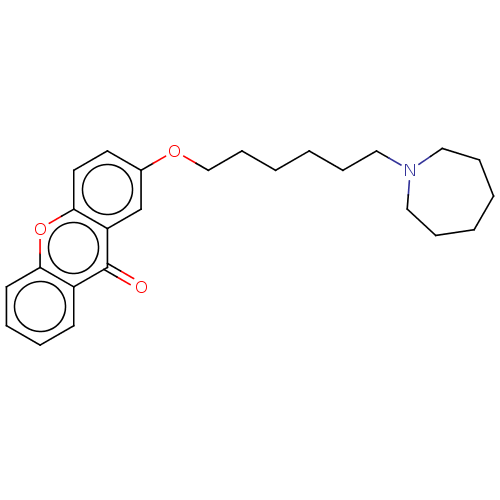
Affinity DataIC50: 8nMAssay Description:Inhibition of recombinant human MAO-B expressed in baculovirus infected BTI cells using p-tyramine as substrate by fluorometric assayMore data for this Ligand-Target Pair
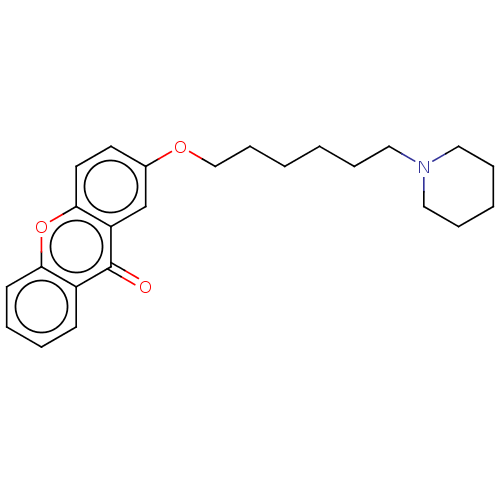
Affinity DataKi: 12nMAssay Description:Displacement of [3H]N-alpha-Methylhistamine from recombinant human histamine H3 receptor expressed in human HEK293 cells incubated for 90 mins by mic...More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] B(Human)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
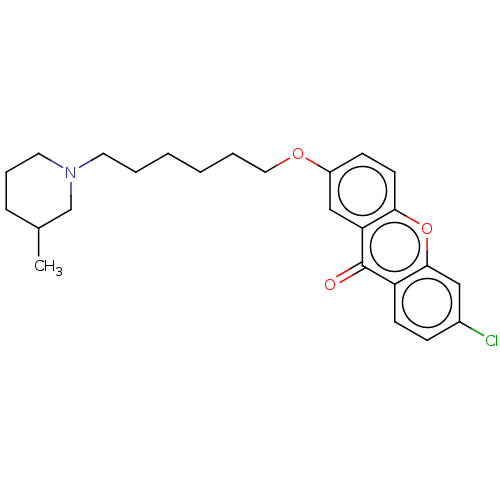
Affinity DataIC50: 25nMAssay Description:Inhibition of recombinant human MAO-B expressed in baculovirus infected BTI cells using p-tyramine as substrate by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of human erythrocyte AChE using ATC iodide as substrate preincubated with enzyme for 4.5 mins followed by substrate addition and measured ...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of human BuChE by spectrophotometric based Ellman's methodMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 67nMAssay Description:Inhibition of Electric eel AChE using ATC iodide as substrate preincubated with enzyme for 4.5 mins followed by substrate addition and measured after...More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Displacement of [3H]-Histamine from human histamine H3 receptor expressed in Sf9 cells incubated for 60 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 86nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 97nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of recombinant human AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectro...More data for this Ligand-Target Pair
Affinity DataIC50: 131nMAssay Description:Inhibition of recombinant human AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectro...More data for this Ligand-Target Pair
Affinity DataIC50: 136nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 148nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 150nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 172nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataKi: 203nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 235nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 238nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataKi: 238nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataKi: 282nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 307nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 308nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 314nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 342nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 353nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 358nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 378nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 384nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] B(Human)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 385nMAssay Description:Inhibition of recombinant human MAO-B expressed in baculovirus infected BTI cells using p-tyramine as substrate by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 386nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 387nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 392nMAssay Description:Inhibition of recombinant human AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectro...More data for this Ligand-Target Pair
Affinity DataIC50: 394nMAssay Description:Inhibition of equine serum BChE using BTCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 406nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataIC50: 421nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate incubated for 5 mins followed by substrate addition and measured after 5 mins by spectrophoto...More data for this Ligand-Target Pair
Affinity DataKi: 436nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataKi: 448nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Inhibition of Electric eel AChE using ATCI as substrate preincubated with enzyme for 60 mins followed by substrate addition and measured for 120 secs...More data for this Ligand-Target Pair
Affinity DataKi: 465nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in CHO-K1 cells by microbeta scintillation analysi...More data for this Ligand-Target Pair
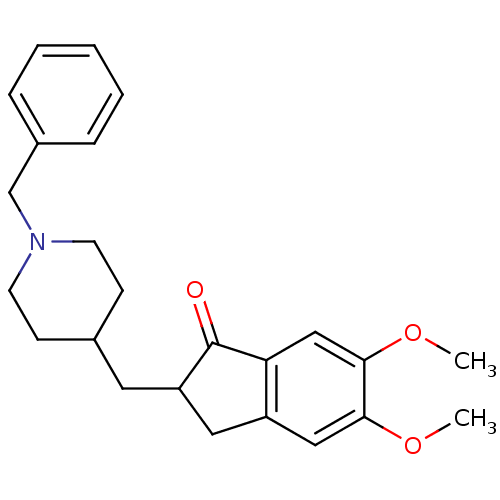
 3D Structure (crystal)
3D Structure (crystal)