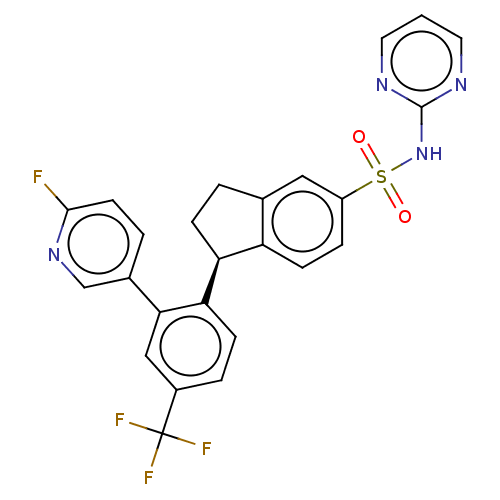
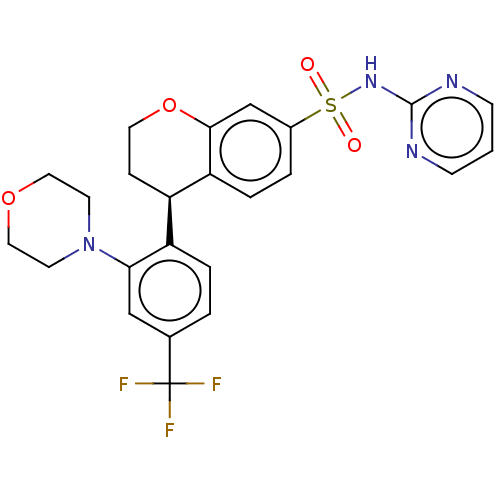
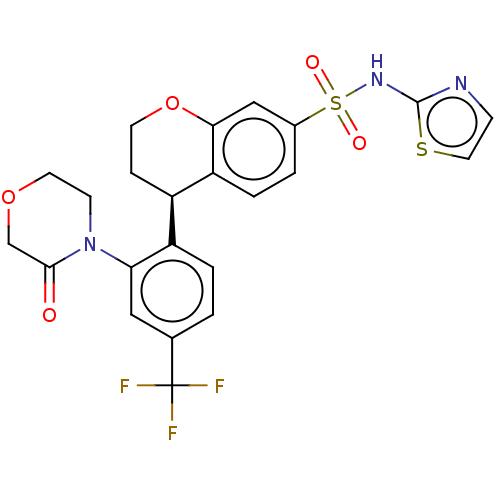
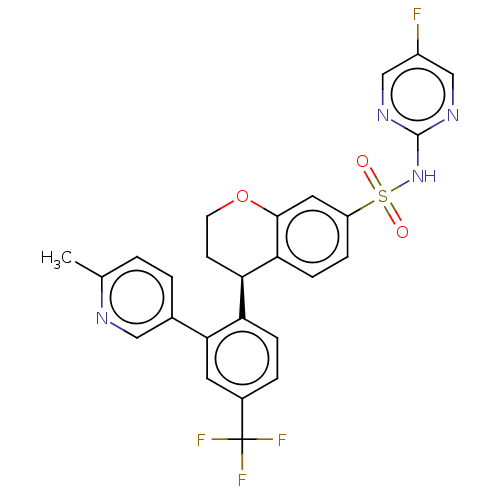
Report error Found 99 Enz. Inhib. hit(s) with all data for entry = 50011193
Affinity DataIC50: 0.110nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 6.5nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 7.20nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of human Nav1.7 expressed in HEK293 cells assessed as V1/2 of slow inactivation at 10 Hz frequency at -120 mV holding potential by single ...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of human Nav1.7 expressed in HEK293 cells assessed as V1/2 of slow inactivation at 10 Hz frequency at -120 mV holding potential by single ...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Use-dependent inhibition of human Nav1.7 expressed in HEK293 cells at 25 pulse 10 Hz frequency at -120 mV holding potential by manual patch clamp ass...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of mouse Nav1.7 expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20 mins follo...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of mouse Nav1.7 expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20 mins follo...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of human Nav1.7 expressed in HEK293 cells assessed as V1/2 of slow inactivation at 10 Hz frequency at -120 mV holding potential by single ...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of mouse Nav1.7 expressed in HEK293 cells assessed as V1/2 of slow inactivation at 10 Hz frequency at -120 mV holding potential by single ...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Use-dependent inhibition of human Nav1.7 expressed in HEK293 cells at 25 pulse 10 Hz frequency at -120 mV holding potential by manual patch clamp ass...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Inhibition of human Nav1.7 expressed in HEK293 cells assessed as V1/2 of slow inactivation at 10 Hz frequency at -120 mV holding potential by single ...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of Nav1.7 (unknown origin) expressed in HEK293 cells assessed as reduction in veratridine-induced depolarization preincubated for 15 to 20...More data for this Ligand-Target Pair