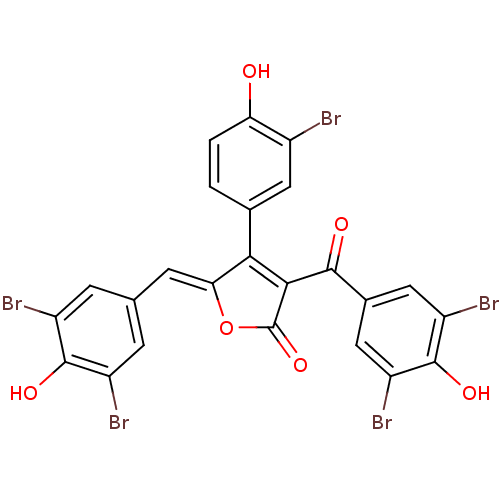
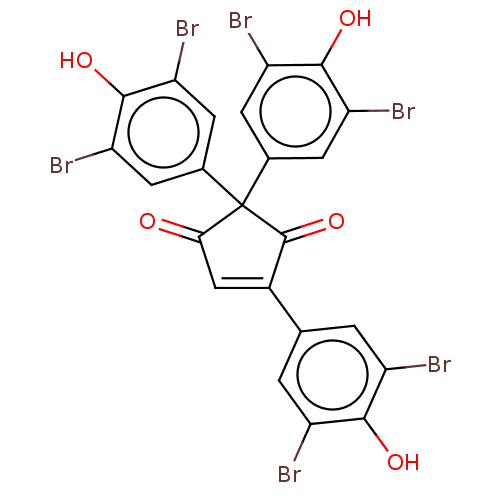
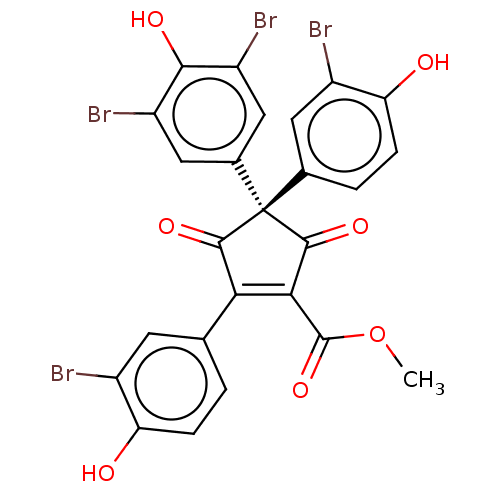
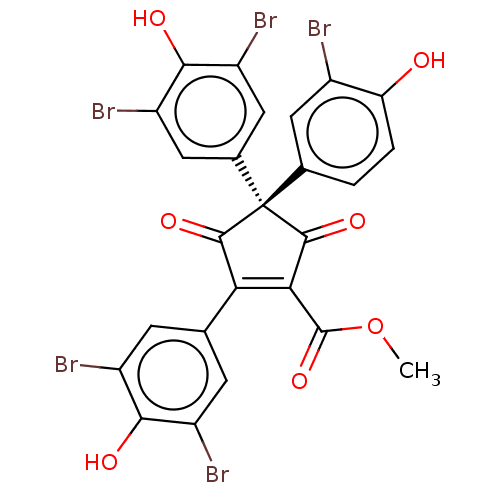
Report error Found 36 Enz. Inhib. hit(s) with all data for entry = 50010350
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90E+4nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+4nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+4nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+4nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.27E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.33E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+5nMAssay Description:Inhibition of Candida albicans ICL assessed as reduction in glyoxylate phenylhydrazone formation using M threo-DL(+)isocitrate as substrate incubated...More data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+5nMAssay Description:Inhibition of Staphylococcus aureus sortase AMore data for this Ligand-Target Pair