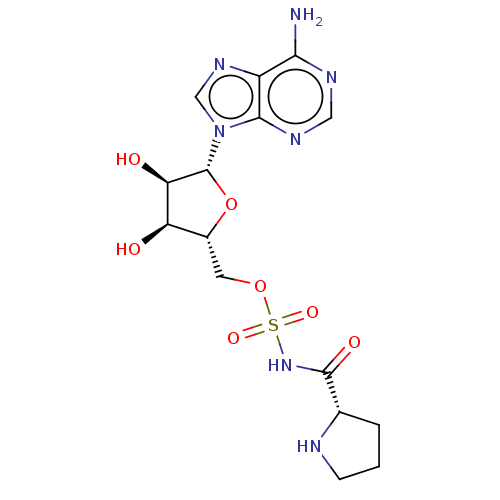
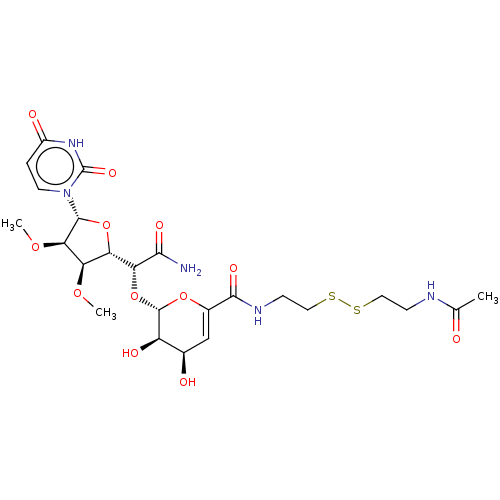
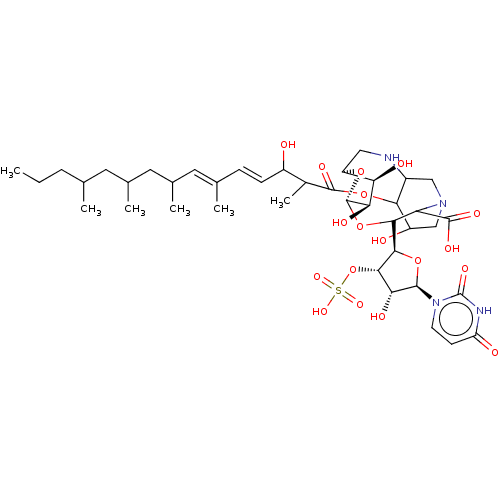
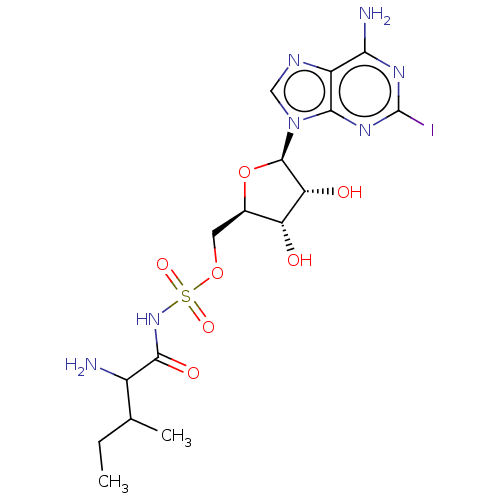
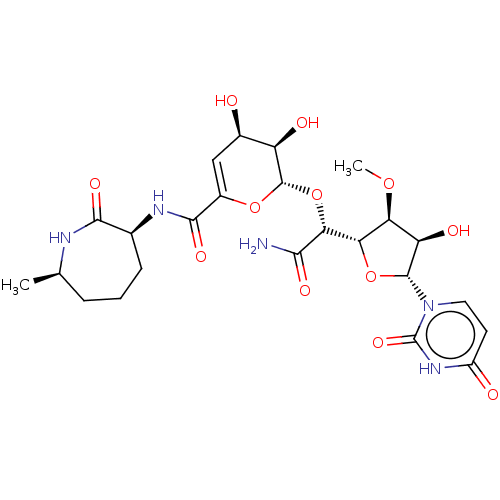
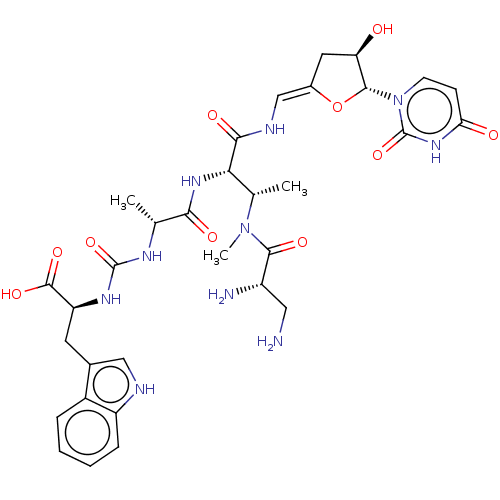
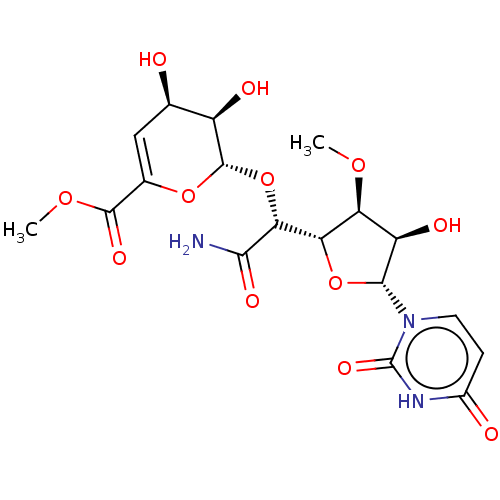
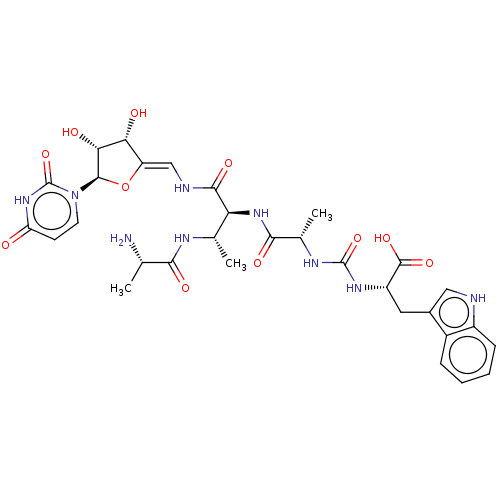
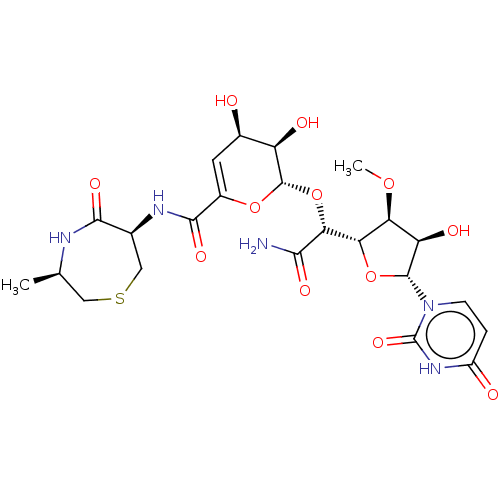
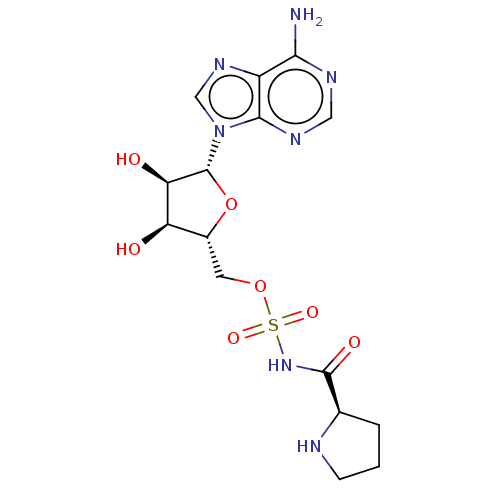
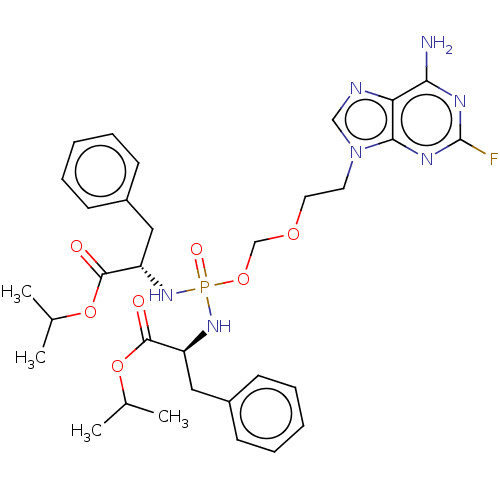
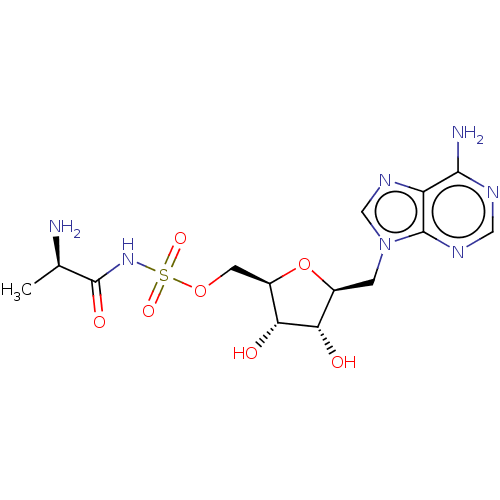
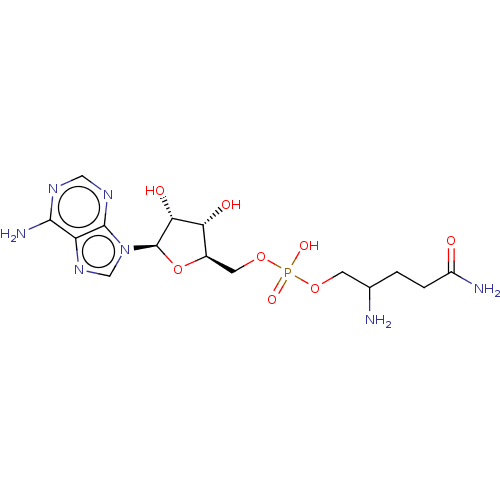
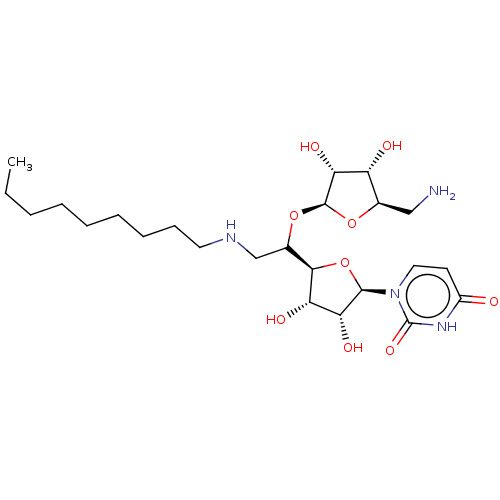
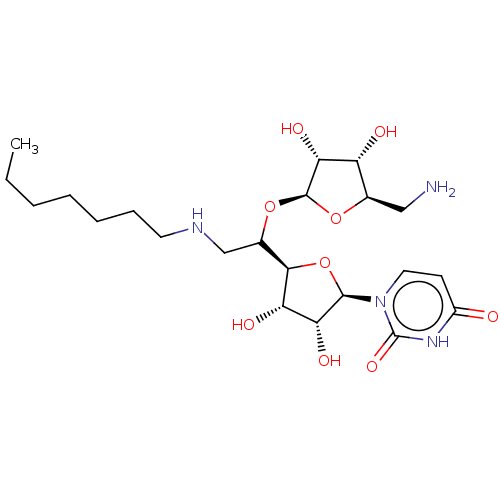
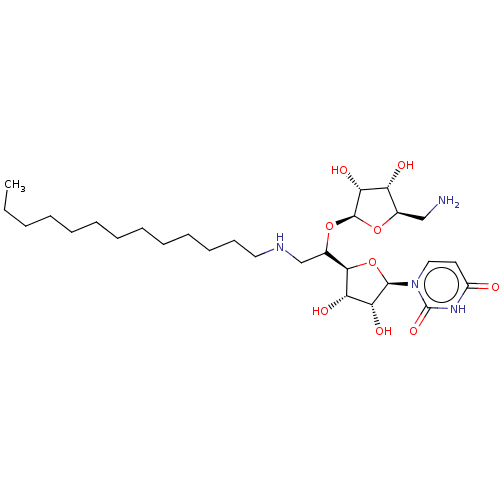
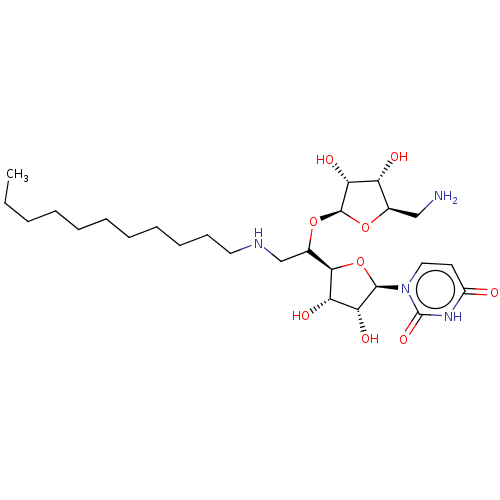
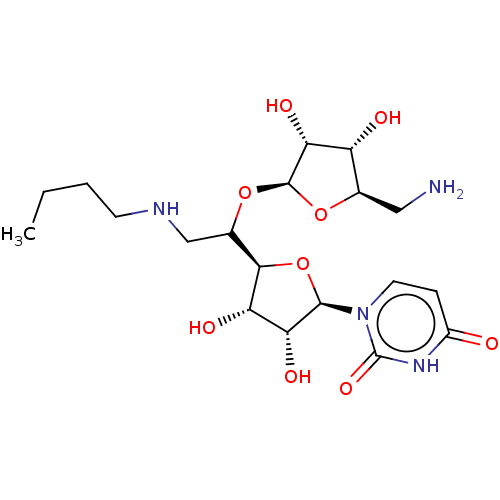
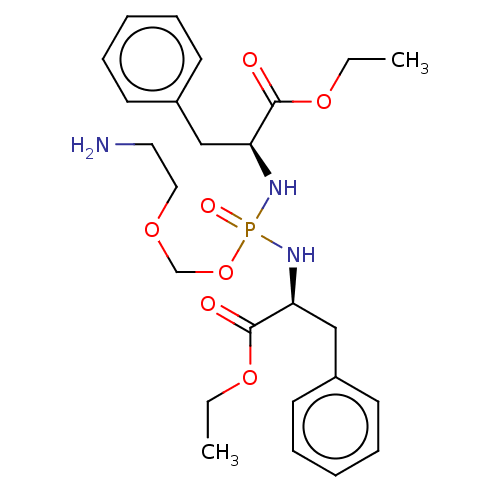
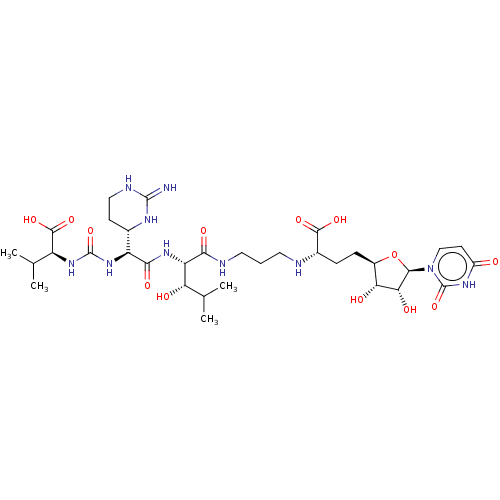
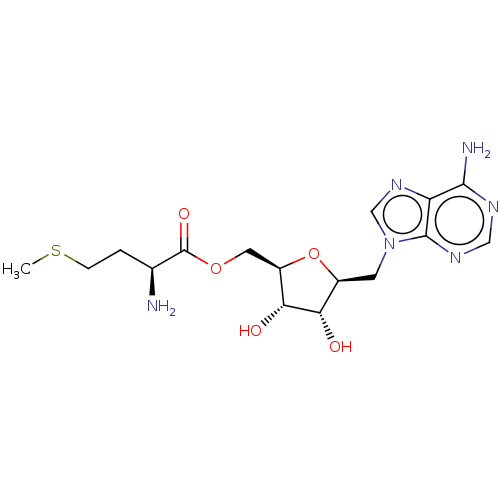
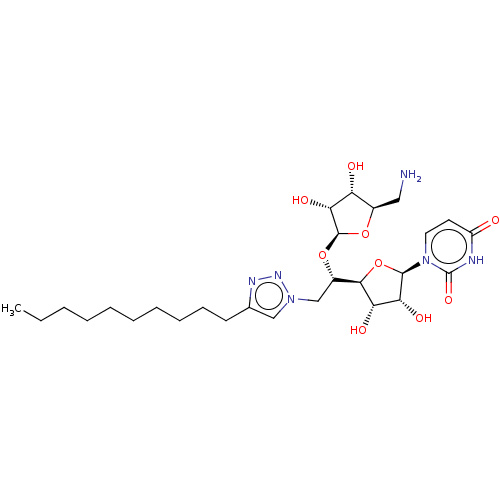
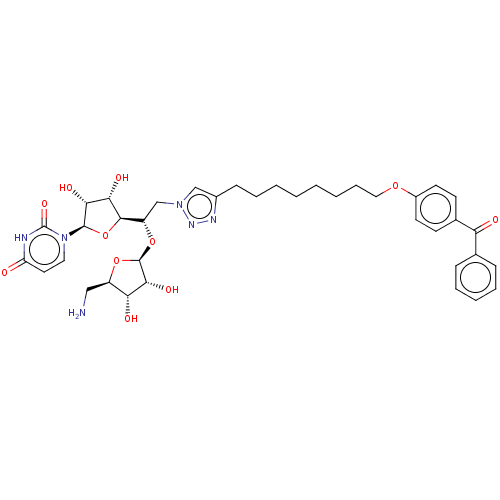
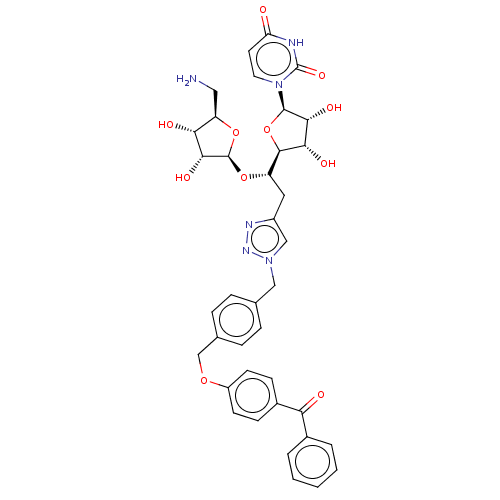
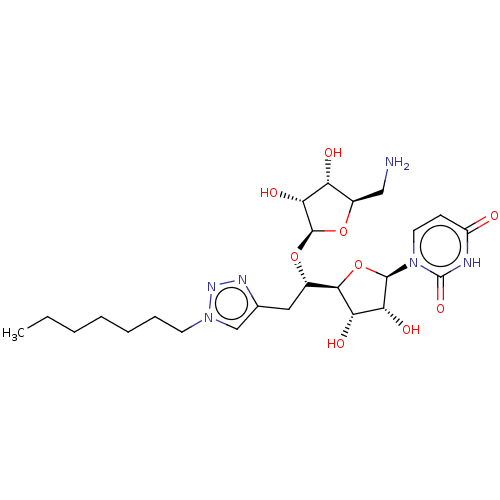
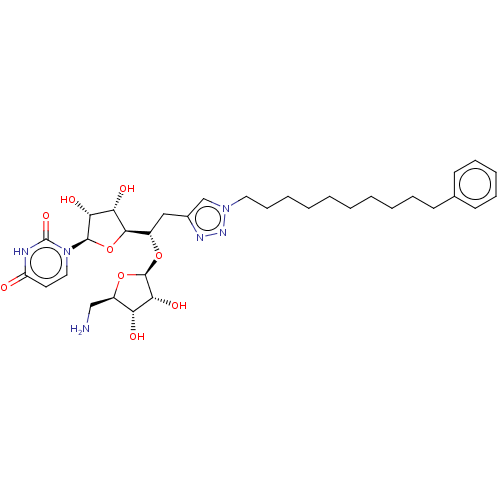
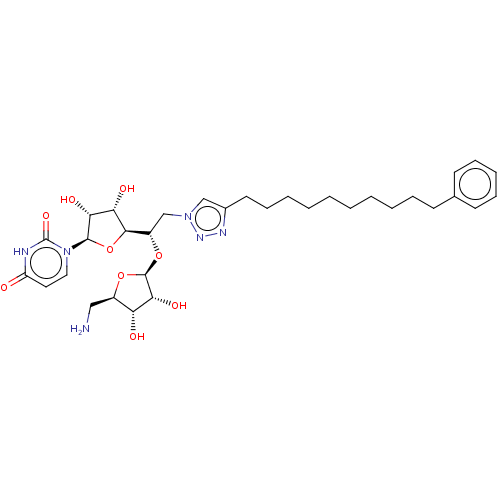
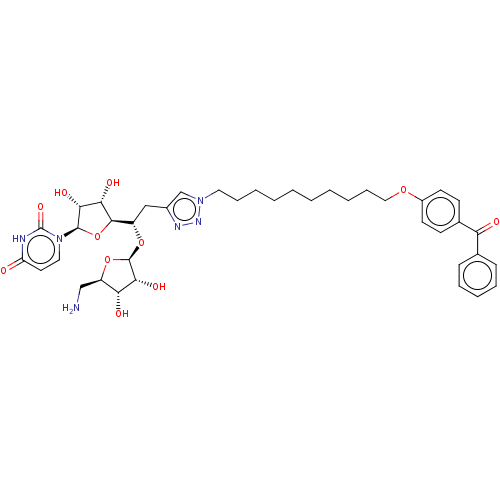
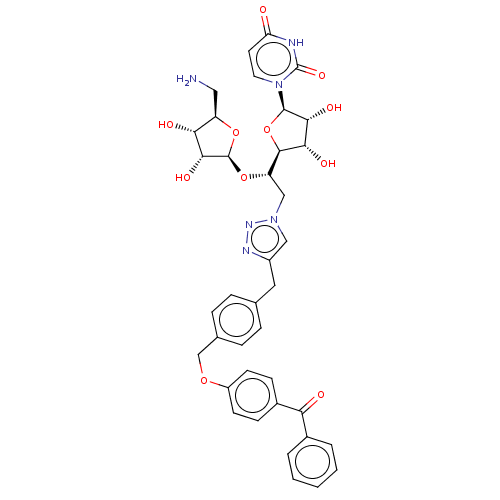
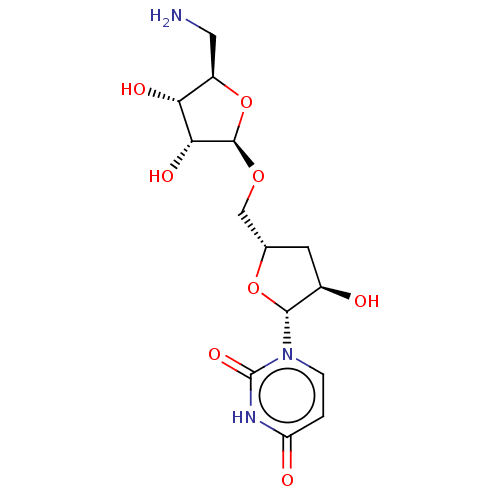
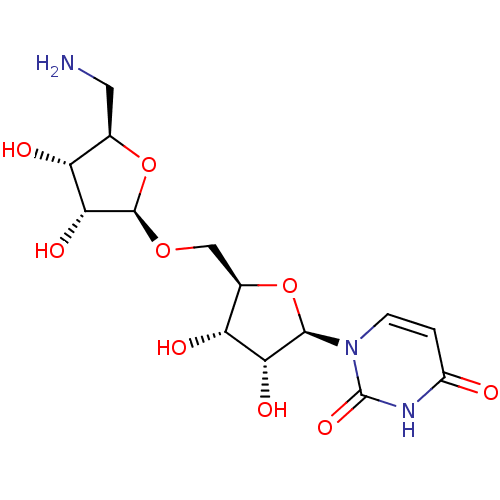
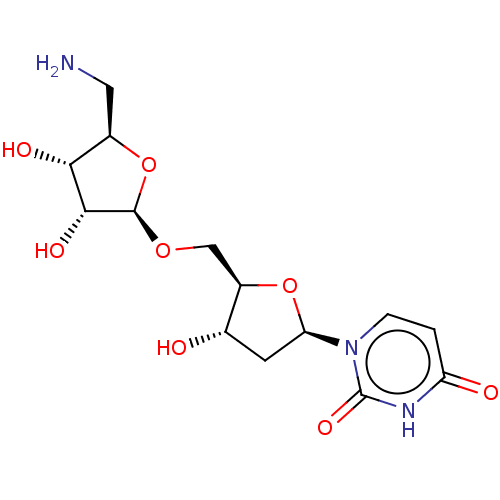
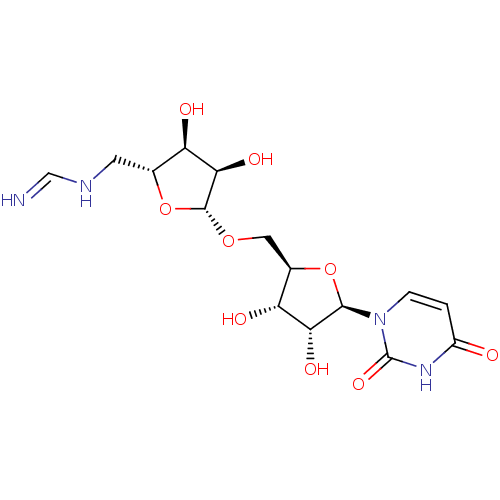
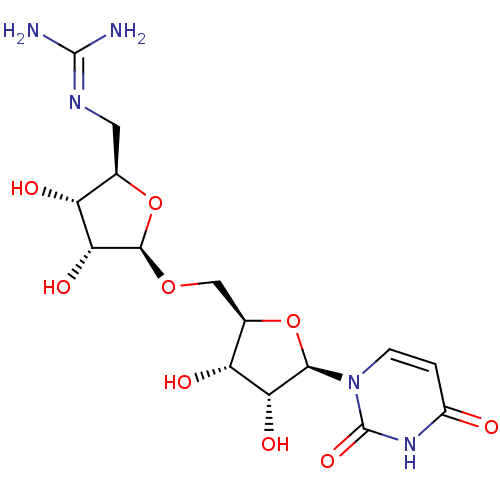
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50018985
Affinity DataKi: 0.600nMAssay Description:Binding affinity to human Prolyl-tRNA synthetaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Binding affinity to Escherichia coli K12 Prolyl-tRNA synthetaseMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetCholesterol side-chain cleavage enzyme, mitochondrial(Rat)
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Antimicrobial activity against Enterococcus faecalis ATCC 29212 assessed as microbial growth inhibitionMore data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of Escherichia coli K12 methionyl tRNA synthetaseMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of Staphylococcus aureus MraYMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Bacillus subtilis (strain 168))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of Bacillus subtilis MraYMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Inhibition of Staphylococcus aureus MraYMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Bacillus subtilis (strain 168))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 42nMAssay Description:Inhibition of Bacillus subtilis MraYMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 58nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 85nMAssay Description:Binding affinity to human Prolyl-tRNA synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 145nMAssay Description:Inhibition of ACT (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 232nMAssay Description:Binding affinity to ADK (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 280nMAssay Description:Binding affinity to Escherichia coli K12 glutaminyl-tRNA synthetase assessed as inhibition constantMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:Binding affinity to Escherichia coli K12 Prolyl-tRNA synthetaseMore data for this Ligand-Target Pair
Affinity DataKi: 690nMAssay Description:Binding affinity to human HGPRT assessed as inhibition of constantMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of Escherichia coli K12 methionyl tRNA synthetaseMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of MraY in Escherichia coli K12More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Escherichia coli (strain K12))
Cardiff University
Curated by ChEMBL
Cardiff University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of MraY in Escherichia coliMore data for this Ligand-Target Pair