Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 50046060
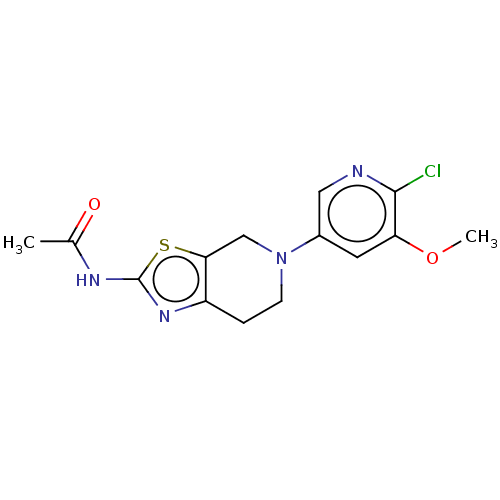
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.00200nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
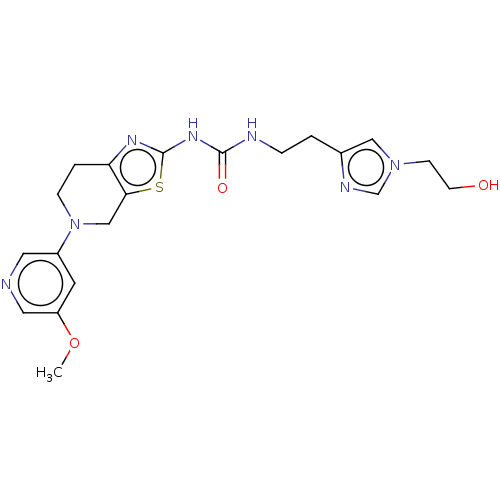
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.00300nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
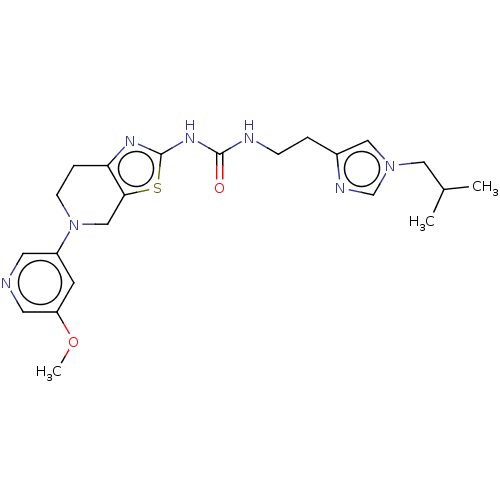
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.00300nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
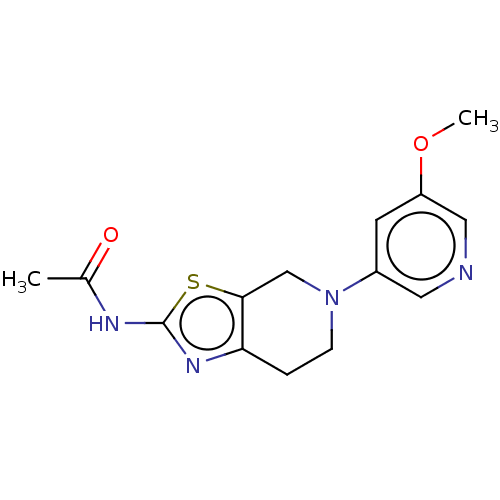
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.00600nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.00900nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0100nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0110nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0130nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0160nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0170nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0240nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0250nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0340nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0480nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.0660nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.157nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 0.330nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using [33P]ATP and PIP2 incubated for 15 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 185nMAssay Description:Inhibition of PI3Kgamma in human THP1 cells assessed as inhibition of MCP1-stimulated chemotaxis pre-incubated for 1 hr before MCP1 stimulationMore data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nMAssay Description:Inhibition of DNAPK (unknown origin) using ATP and PPLSQEAFADLWKKK substrate by radiometric peptide assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Inhibition of JAK2 (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.60E+3nMAssay Description:Inhibition of TRKA (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of GSK3beta (unknown origin) using ATP and GSM peptide substrate by radiometric peptide assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of JAK2 (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of GSK3beta (unknown origin) using ATP and GSM peptide substrate by radiometric peptide assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of TRKA (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of FLT3 (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of FLT3 (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of SRC (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of DNAPK (unknown origin) using ATP and PPLSQEAFADLWKKK substrate by radiometric peptide assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of SRC (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of c-KIT (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of c-KIT (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of KDR (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of KDR (unknown origin) using ATP and polyE4Y substrate by radiometric assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of PI3Kbeta in human THP1 cells assessed as inhibition of CSF1-stimulated chemotaxis pre-incubated for 1 hr before MCP1 stimulationMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of PI3Kalpha in human THP1 cells assessed as inhibition of CSF1-stimulated chemotaxis pre-incubated for 1 hr before MCP1 stimulationMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of PI3Kdelta in human THP1 cells assessed as inhibition of CSF1-stimulated chemotaxis pre-incubated for 1 hr before MCP1 stimulationMore data for this Ligand-Target Pair
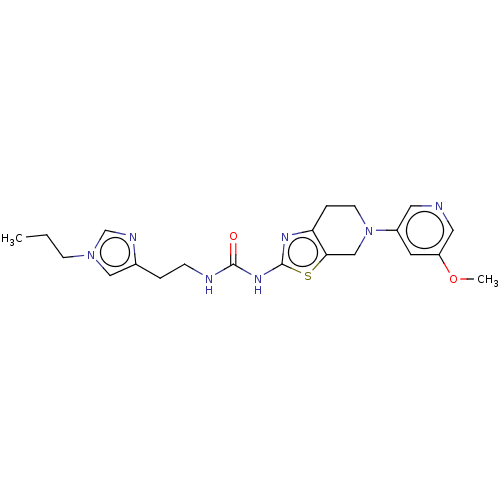
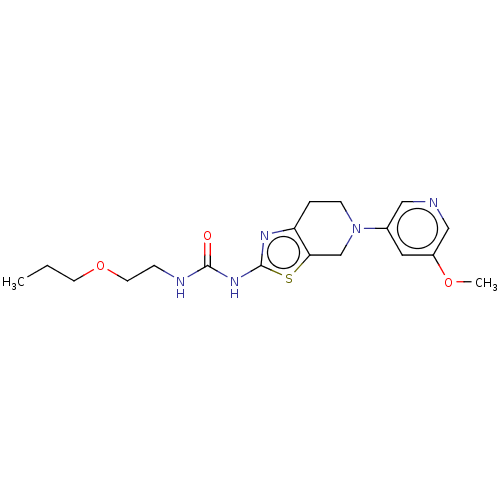
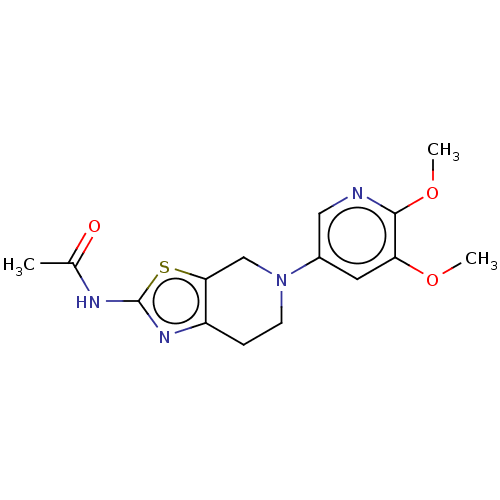
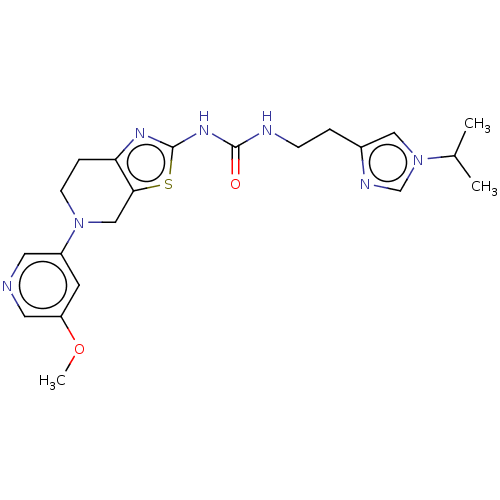
 3D Structure (crystal)
3D Structure (crystal)