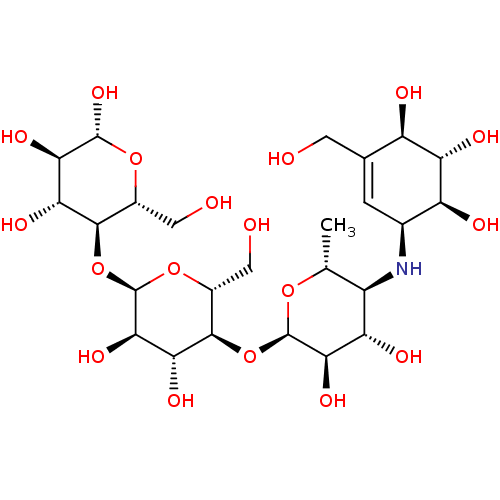
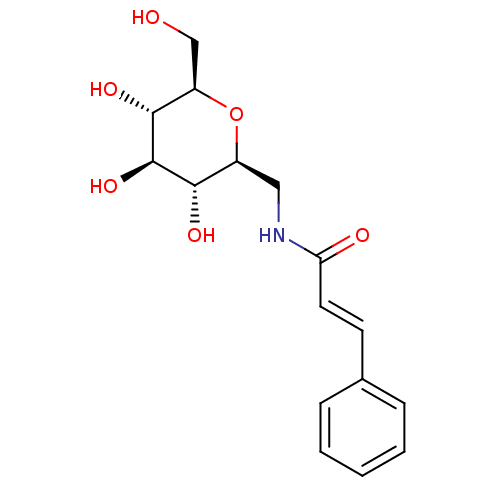
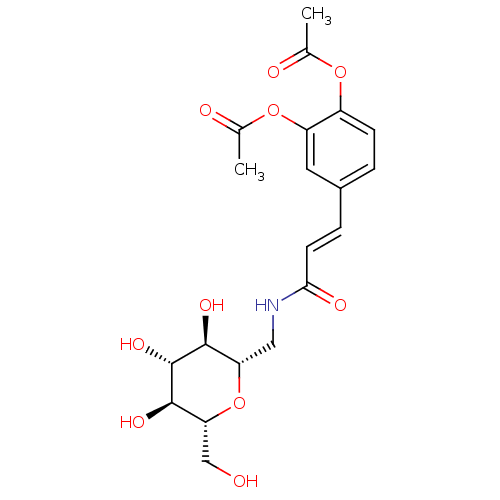
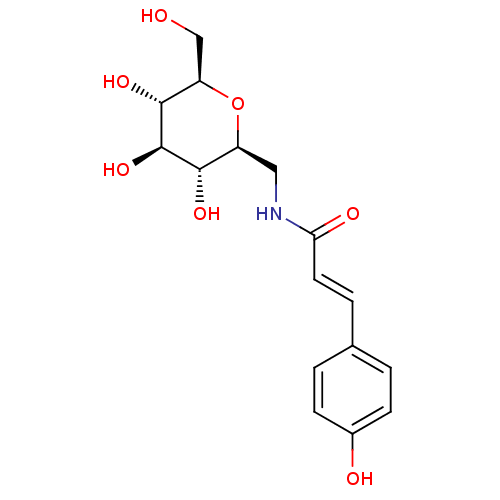
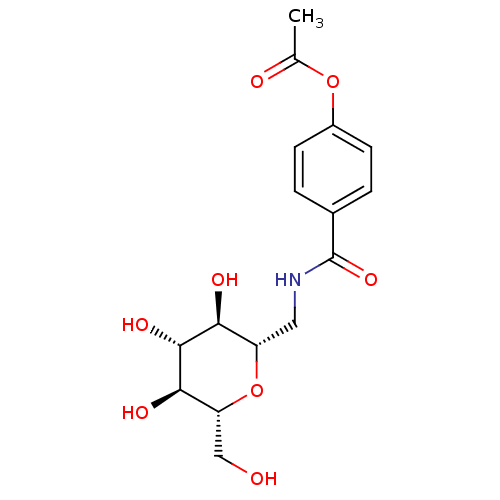
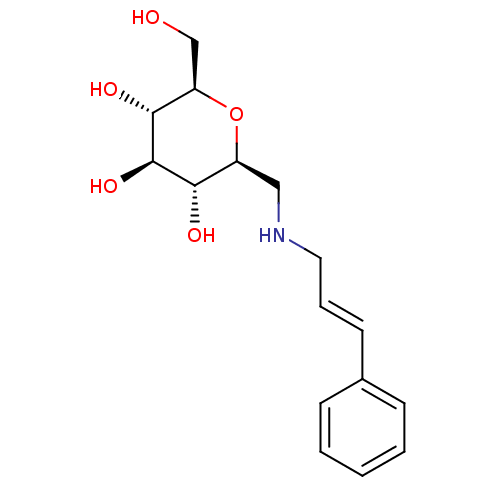
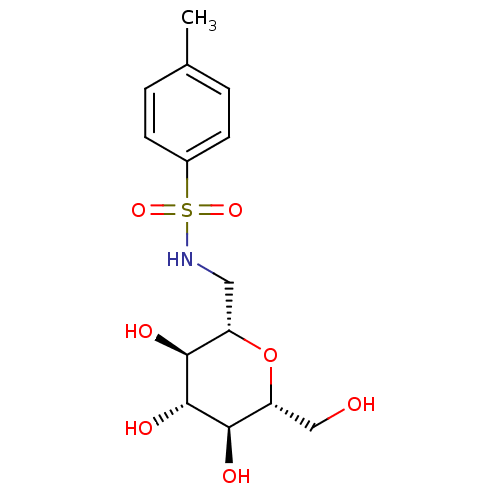
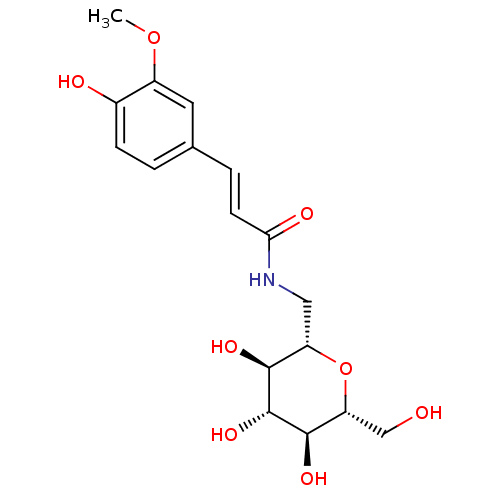
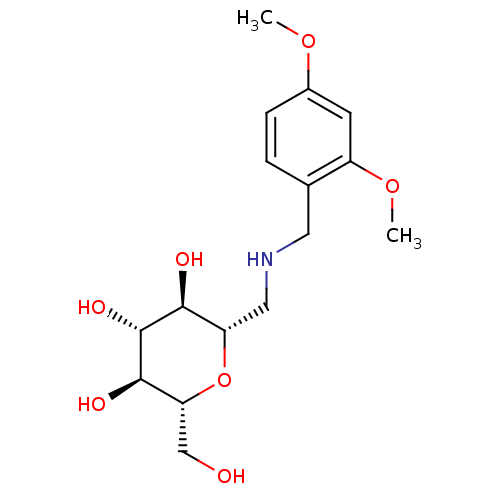
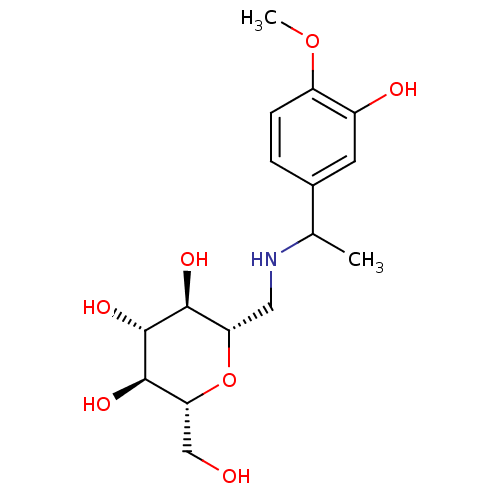
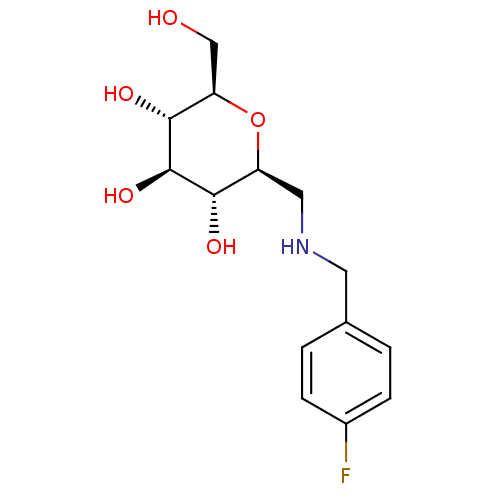
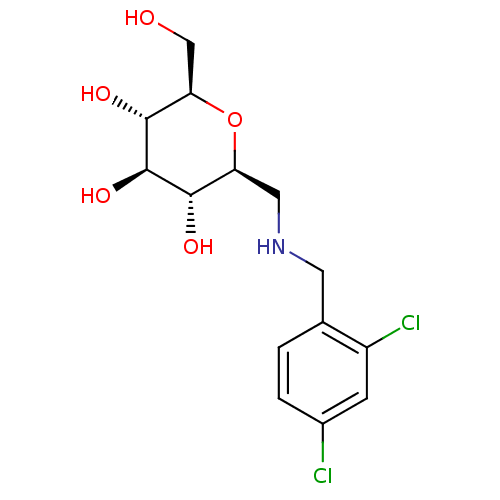
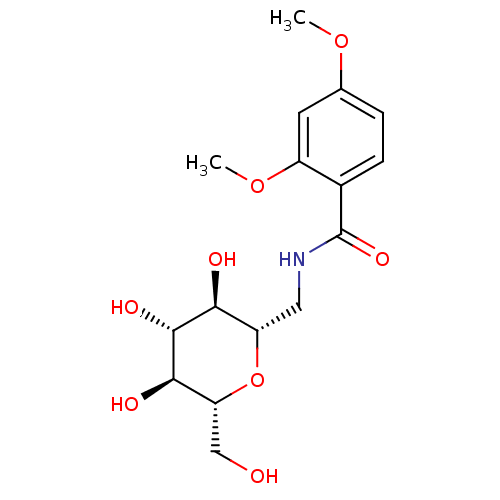
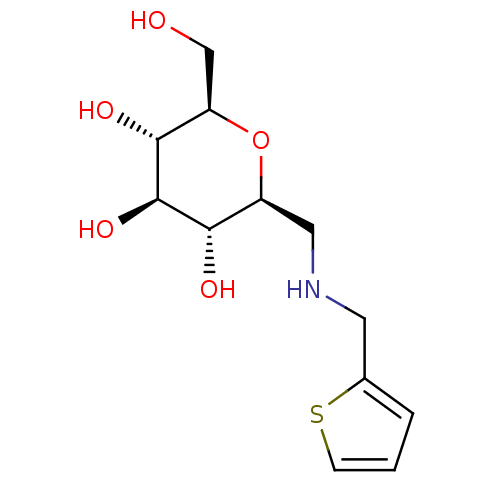
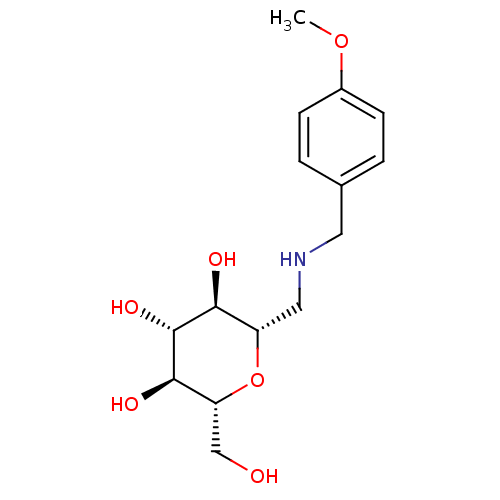
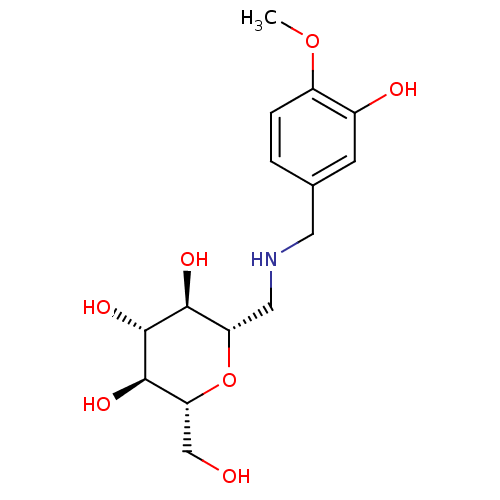
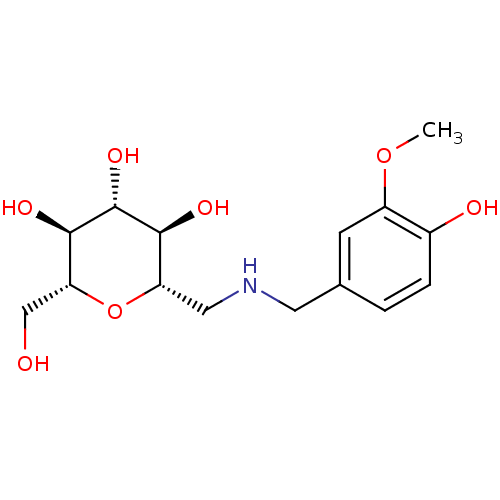
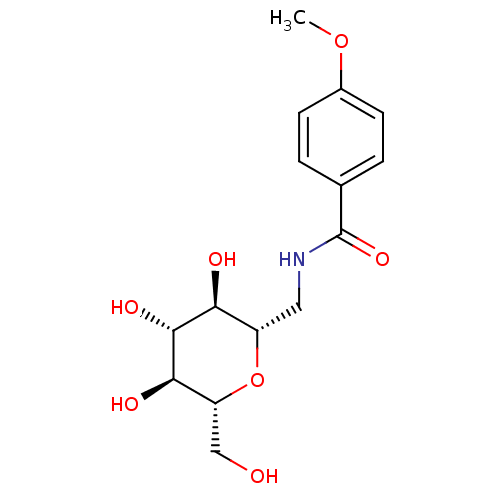
Report error Found 107 Enz. Inhib. hit(s) with all data for entry = 50043270
Affinity DataIC50: 400nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:Competitive inhibition of rat intestinal maltase using maltose as substrate by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60E+3nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+4nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataKi: 1.71E+4nMAssay Description:Competitive inhibition of rat intestinal sucrase using sucrose as substrate by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.62E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 3.92E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 4.43E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 9.65E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 9.74E+4nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.02E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.07E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.49E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.54E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.65E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.68E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.72E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.84E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.86E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.86E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.21E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+5nMAssay Description:Inhibition of rat intestinal maltase using maltose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.25E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 2.31E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as 4-nitrophenol release from 4-nitrophenyl alpha-D-glucopyranoside preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 2.53E+5nMAssay Description:Inhibition of rat intestinal sucrase using sucrose as substrate assessed as D-glucose release from substrate preincubated for 15 mins measured after ...More data for this Ligand-Target Pair