Report error Found 17 Enz. Inhib. hit(s) with all data for entry = 4742
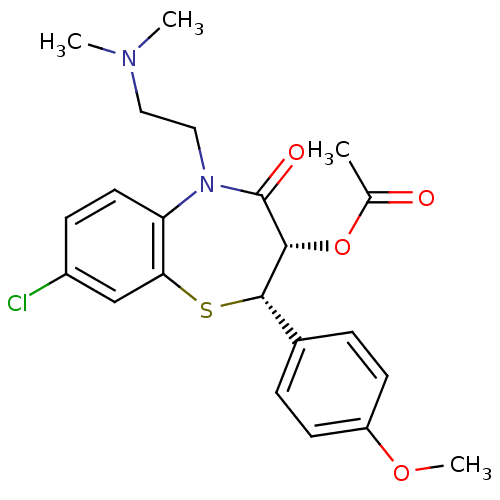
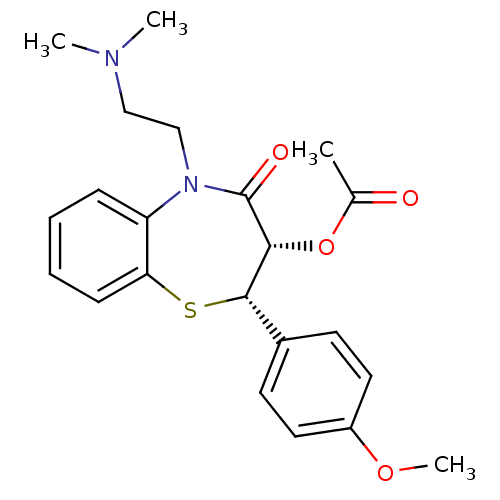
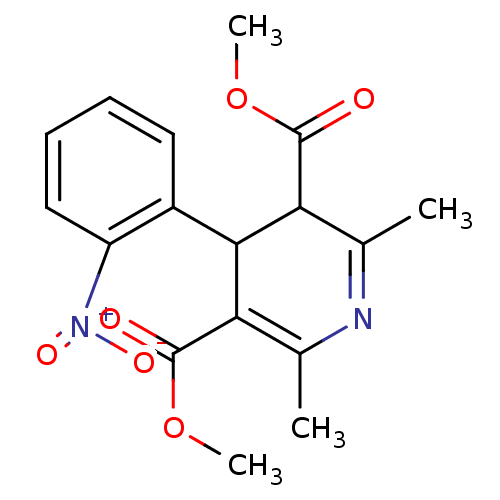
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(Rat)
Marion Merrell Dow
Curated by PDSP Ki Database
Marion Merrell Dow
Curated by PDSP Ki Database