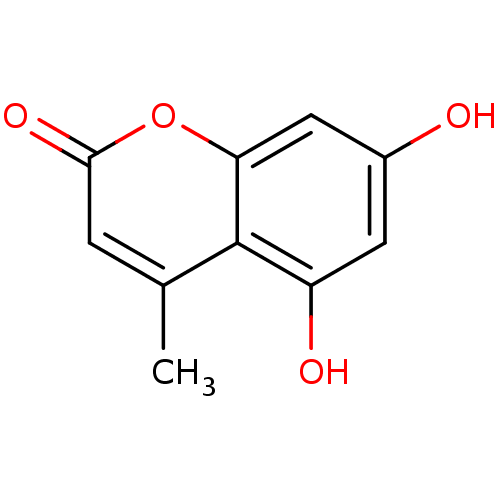
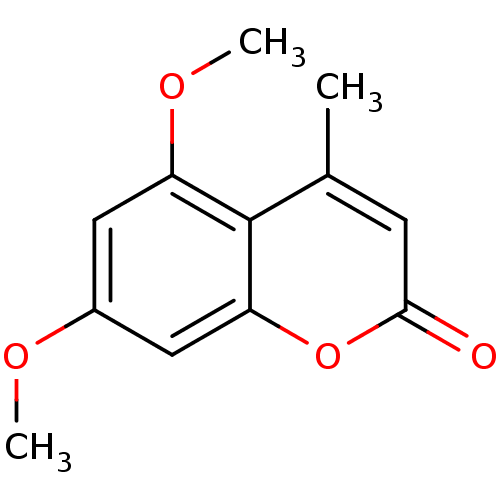
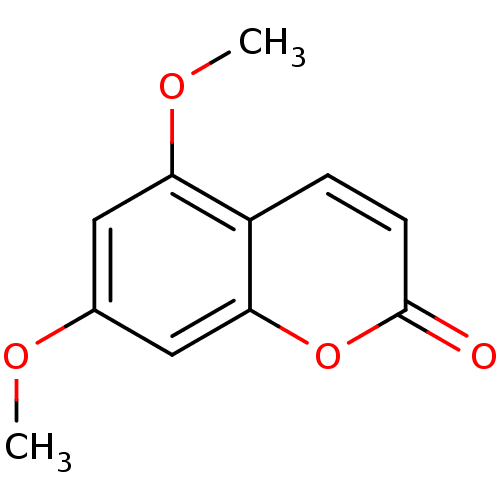
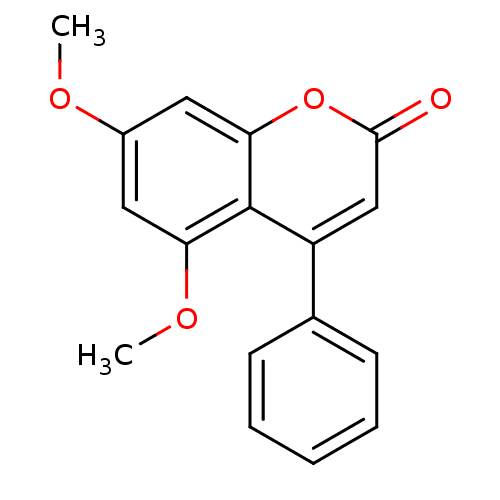
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 5771
Affinity DataIC50: 1.79E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 2.62E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.03E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.38E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.66E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.84E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 3.97E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 4.16E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 4.58E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 4.72E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 5.41E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+4nMpH: 7.0 T: 2°CAssay Description:Inhibition assay using HCV NS5B.More data for this Ligand-Target Pair