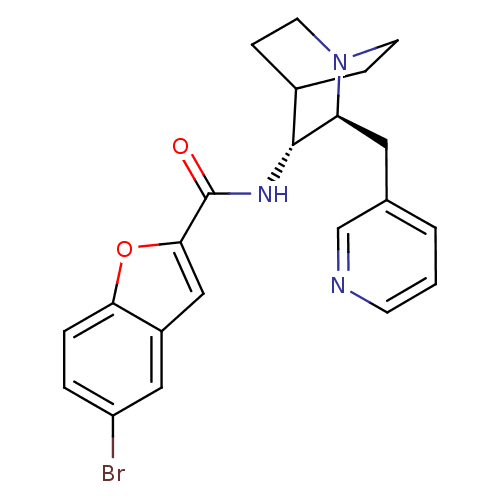
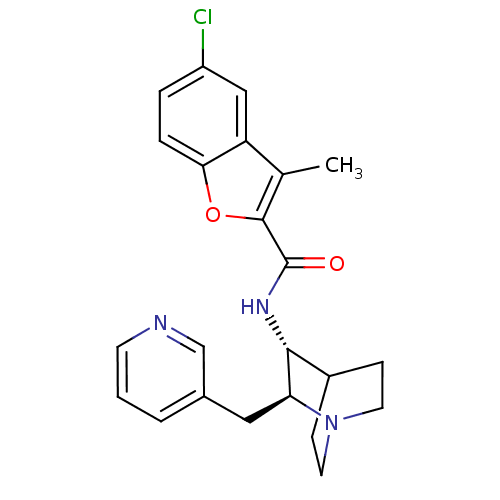
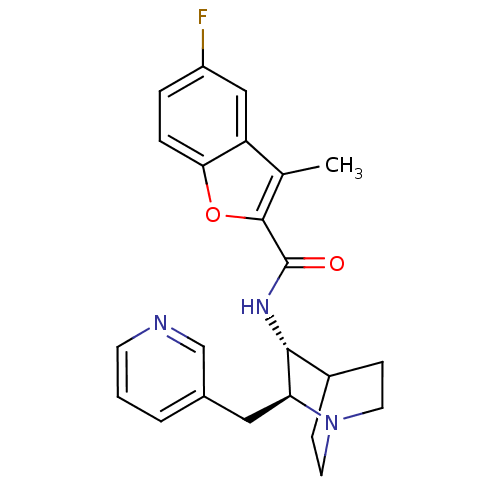
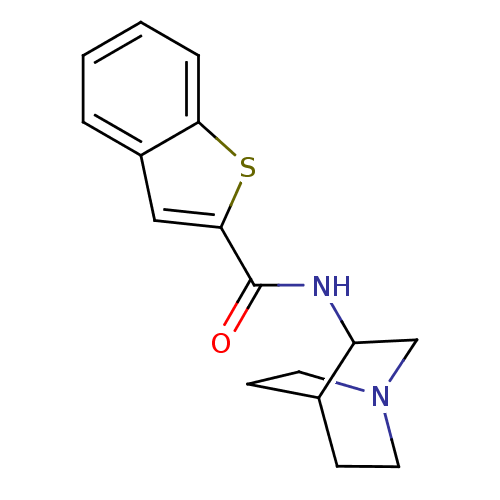
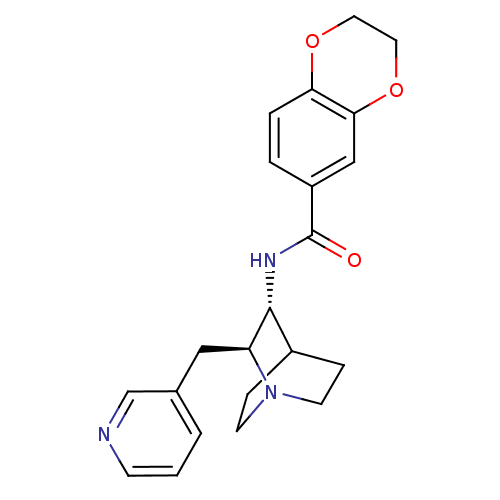
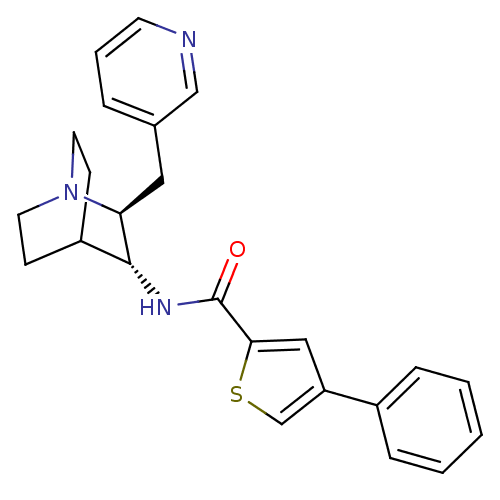
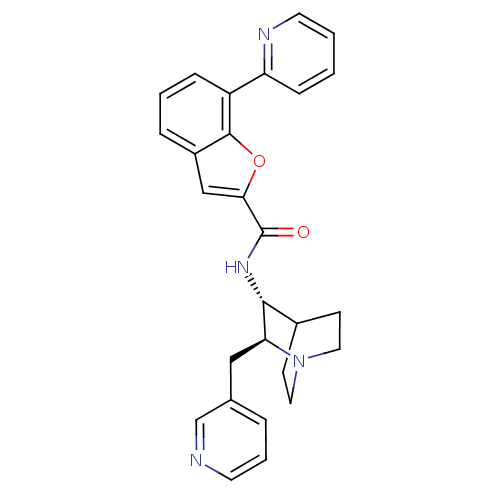
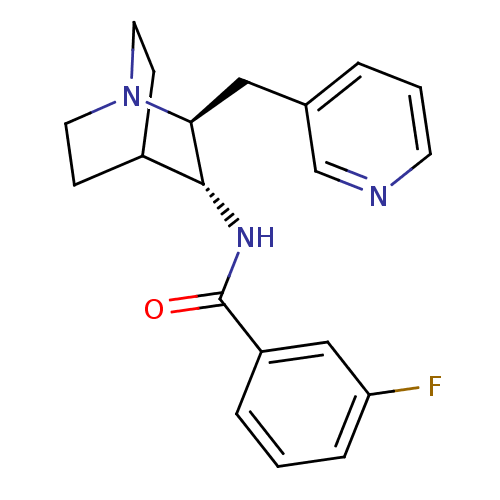
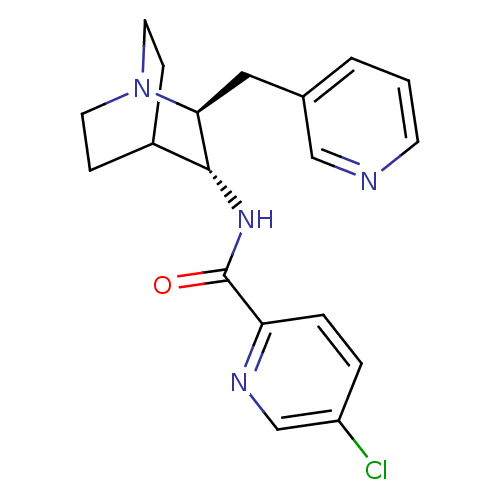
Report error Found 213 Enz. Inhib. hit(s) with all data for entry = 50040768
Affinity DataKi: 0.420nMAssay Description:Displacement of [3H]methyllycaconitine form alpha7 nAchR in rat hippocampal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.510nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 1nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]nicotine human alpha4beta2 nAChR in SH-EP1 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]nicotine human alpha4beta2 nAChR in SH-EP1 cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.70nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.98nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.20nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.20nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 9.98nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 11nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 11.2nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 11.5nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 12nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 12.3nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 14nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataEC50: 14nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 14.6nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 15.7nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 17nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 22.1nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 24.1nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 26.3nMAssay Description:Displacement of [3H]methyllycaconitine form alpha7 nAchR in rat hippocampal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 29.1nMAssay Description:Displacement of [3H]methyllycaconitine form alpha7 nAchR in rat hippocampal membranesMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Agonist activity at rat alpha7 nAchR expressed in GH4C1 cells by whole cell patch clamp assayMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]epibatidine form human alpha7 nAchR expressed in HEK293 cellsMore data for this Ligand-Target Pair
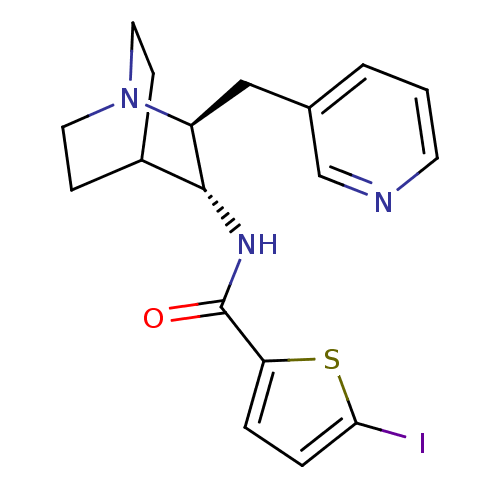
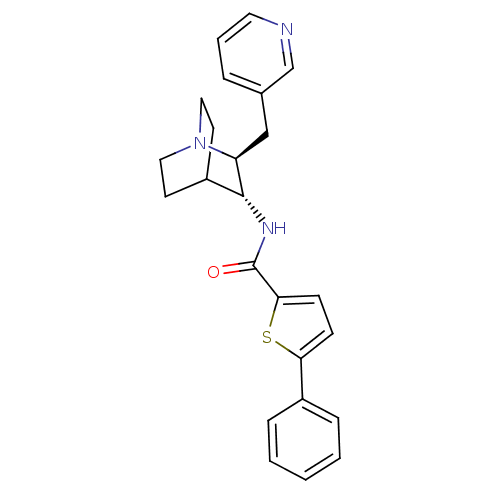
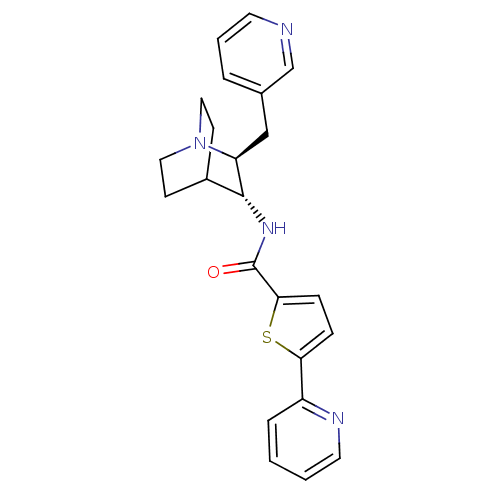
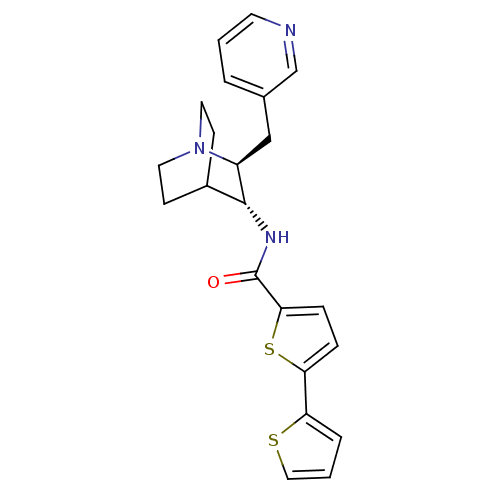
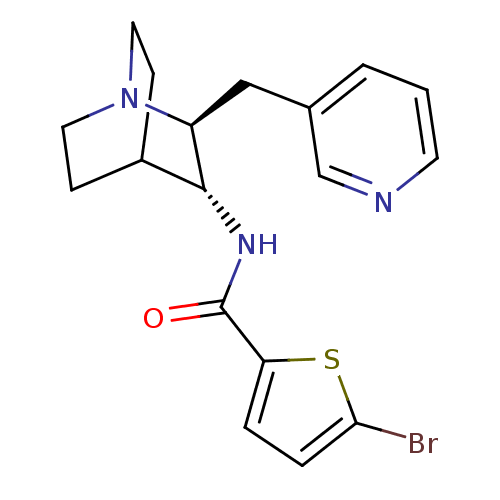
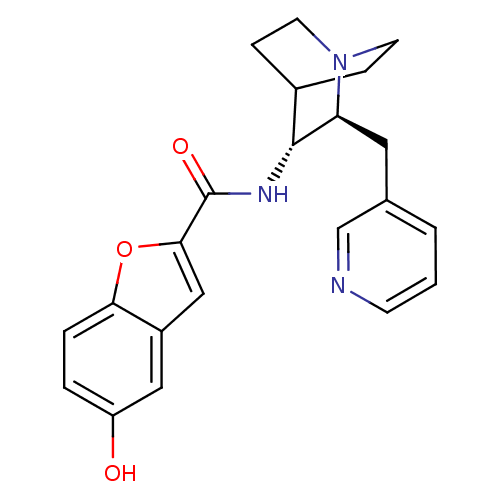
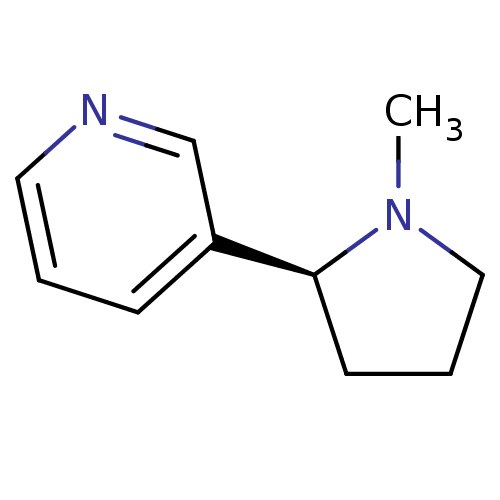
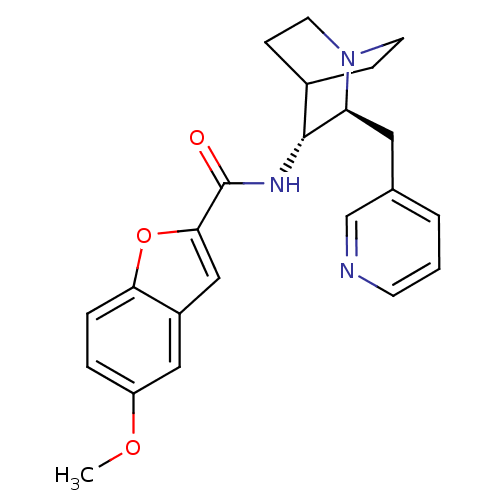
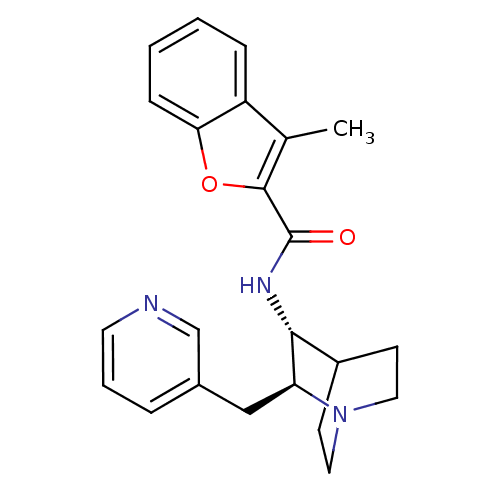
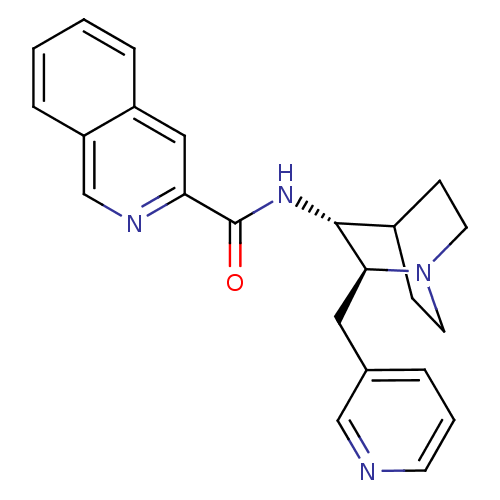
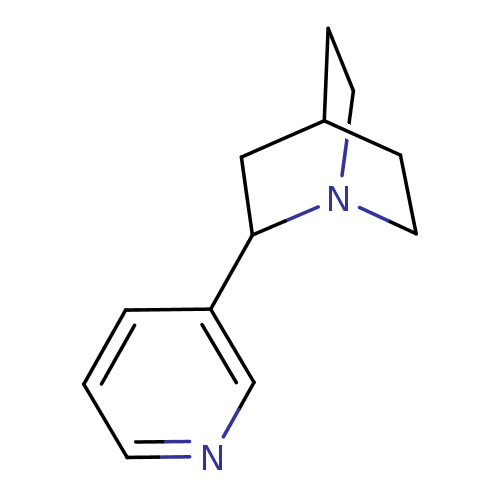
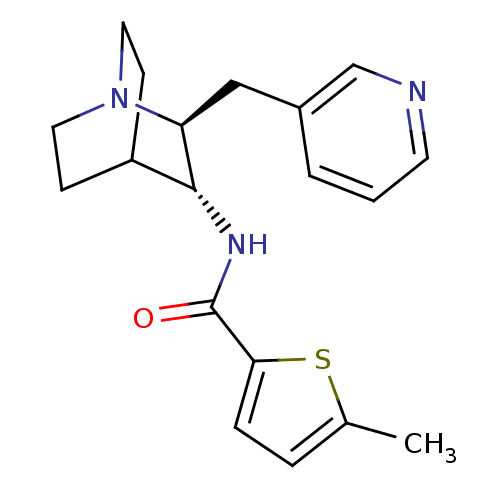
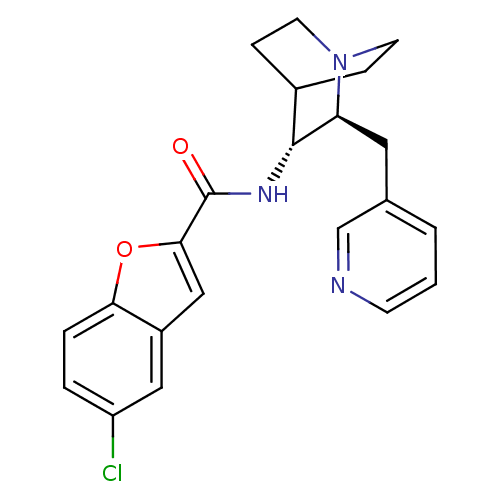
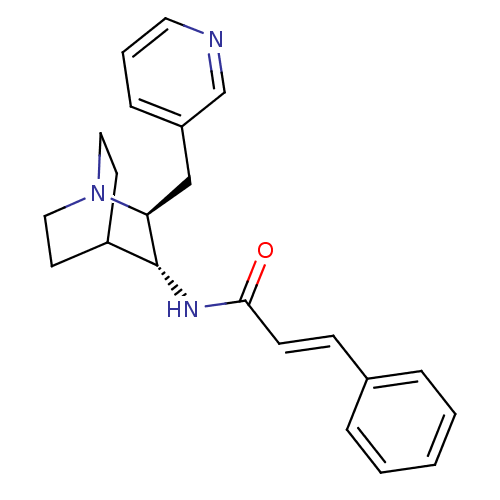
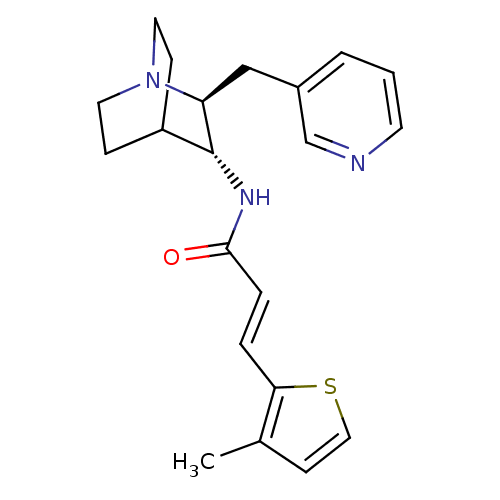
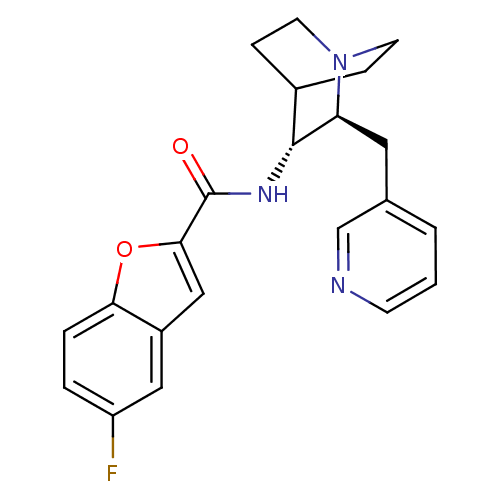
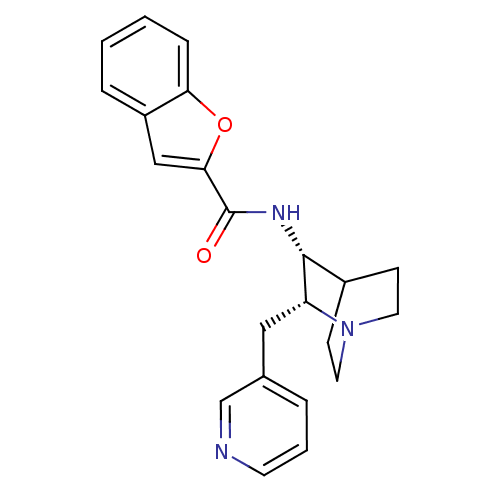
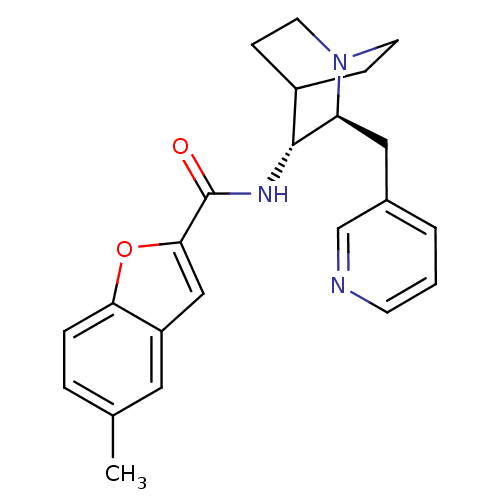
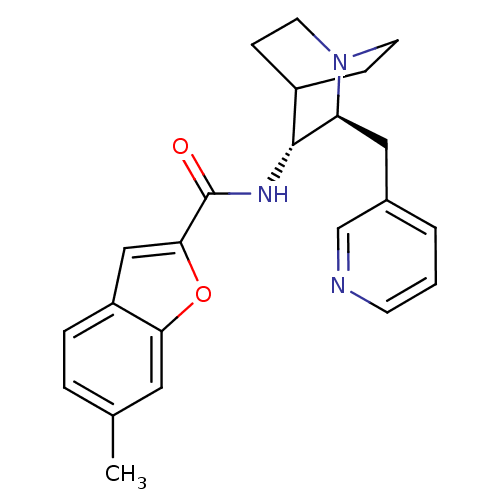
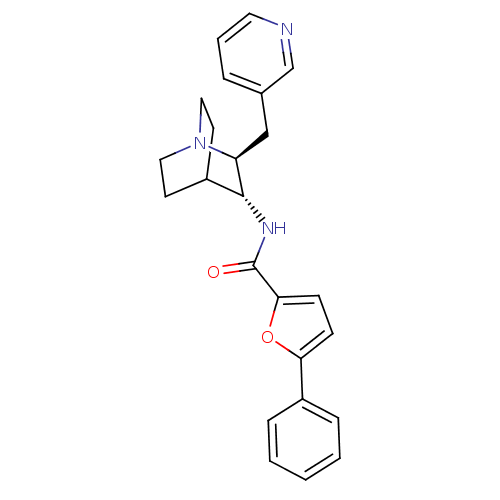
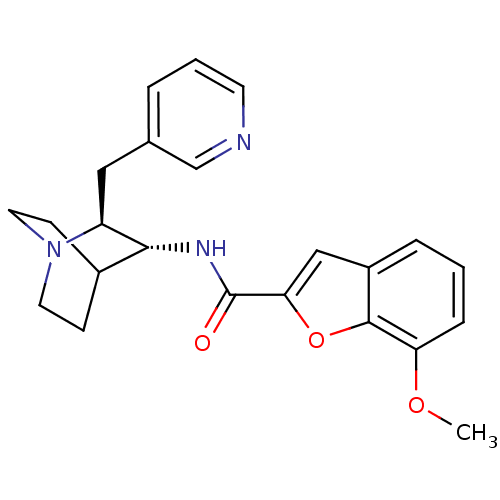
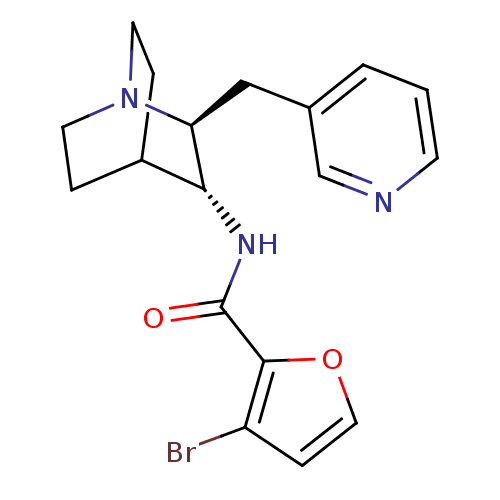

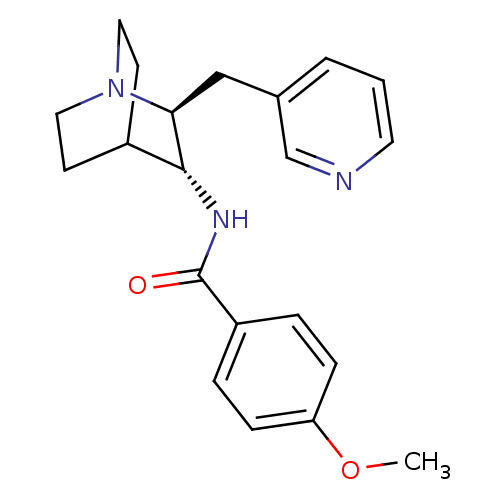
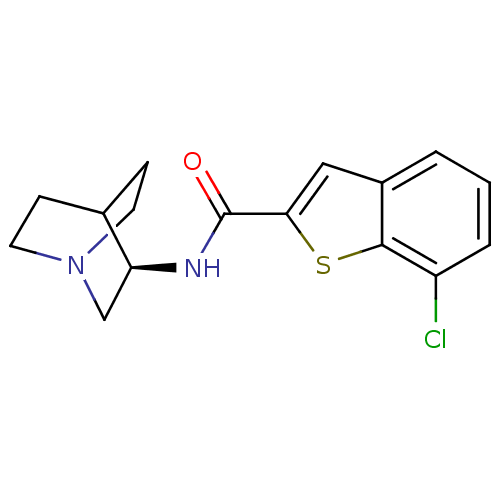
 3D Structure (crystal)
3D Structure (crystal)