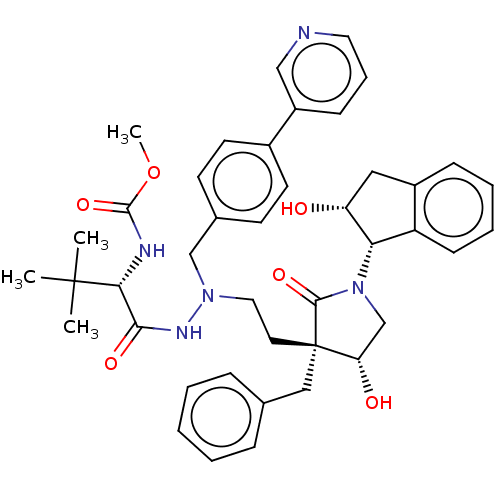
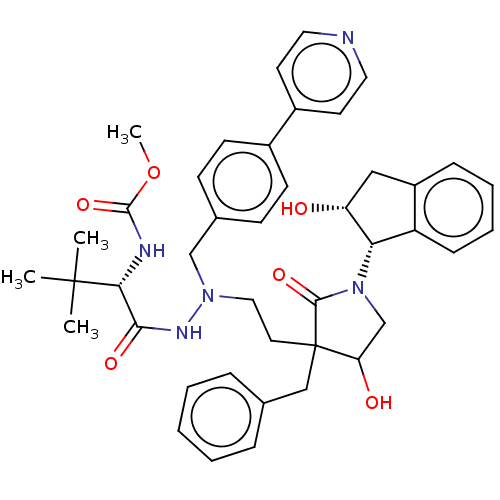
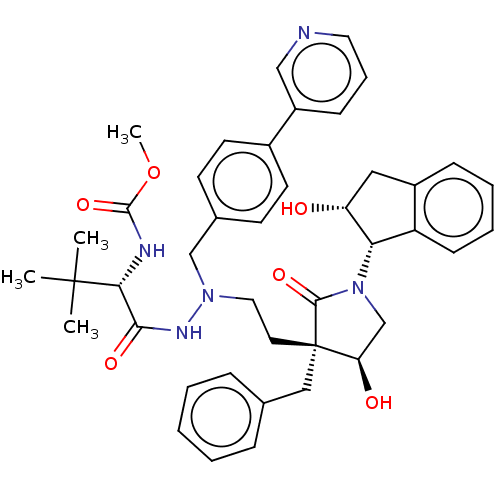
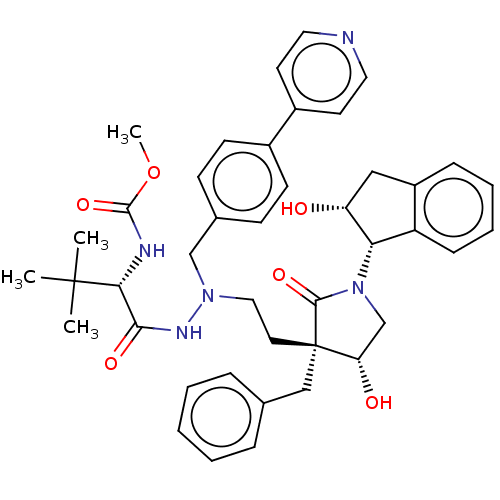
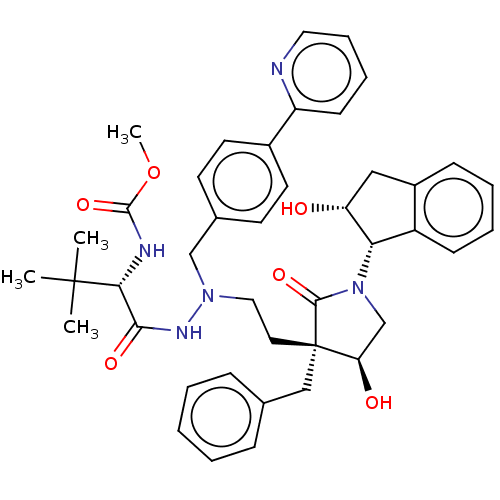
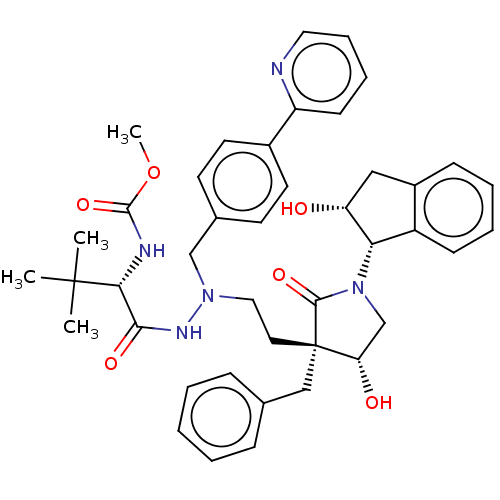
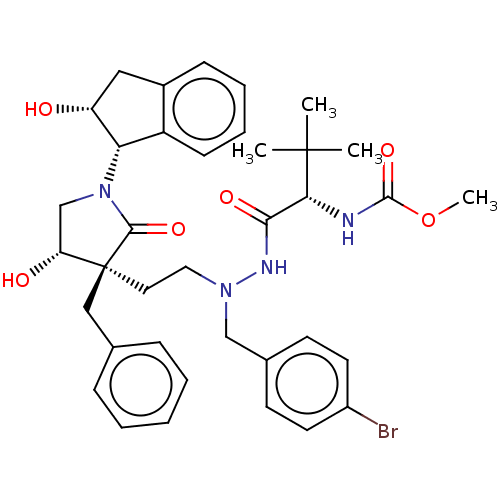
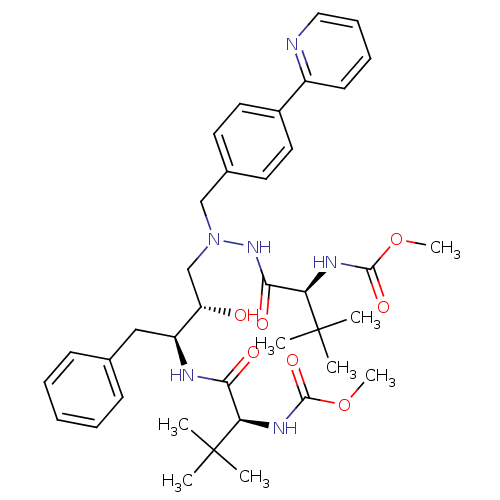
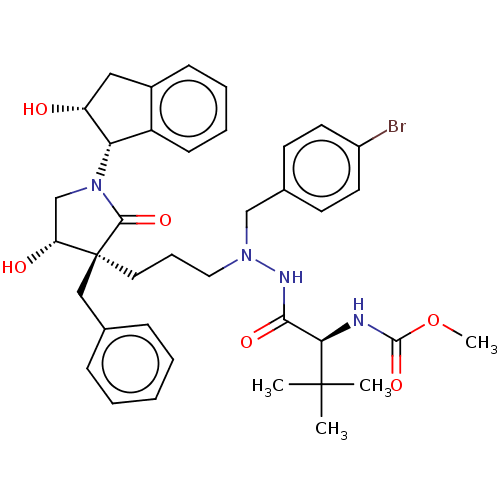
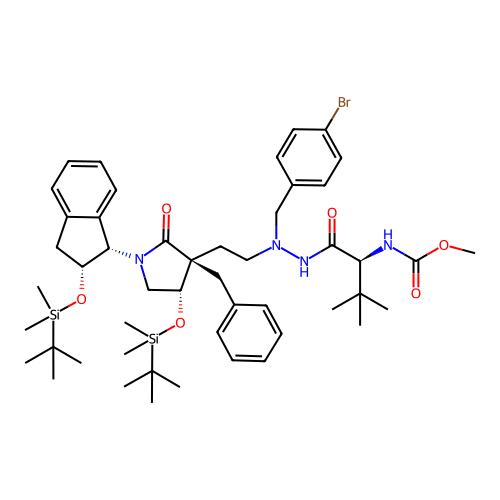
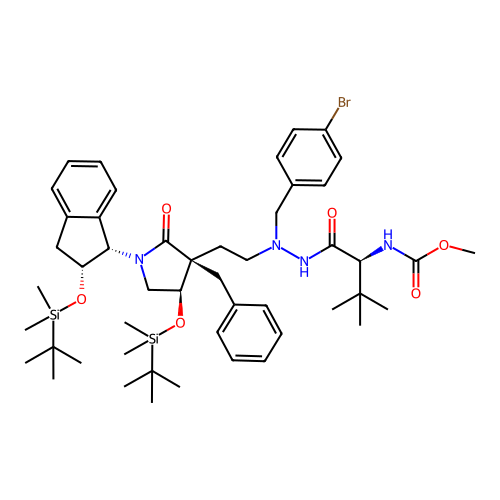
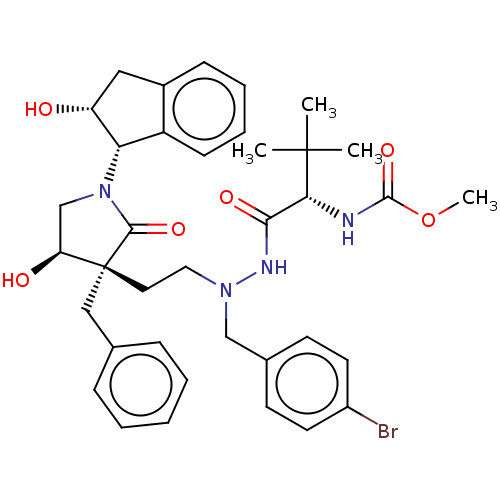
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 50004638
Affinity DataKi: 0.400nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 2.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 760nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 940nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair