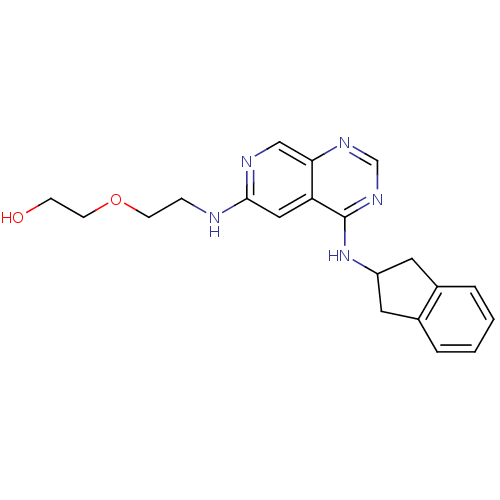
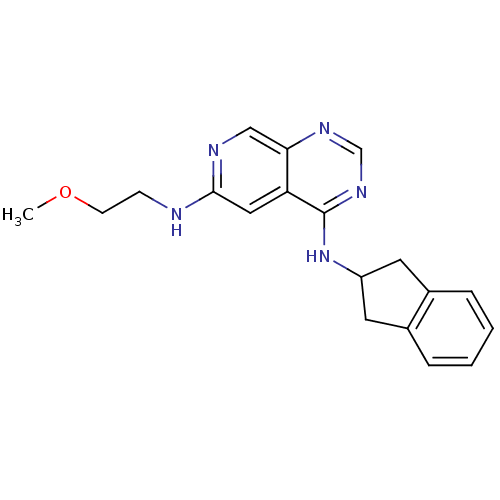
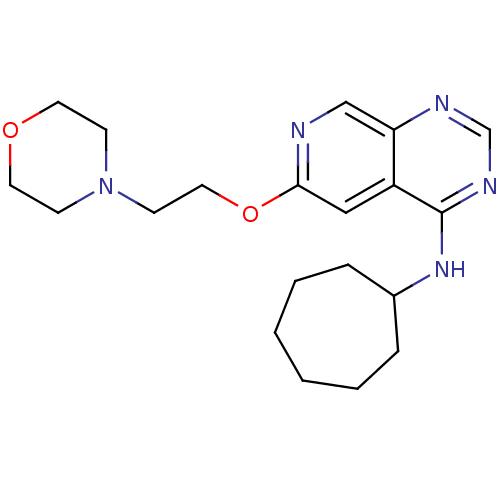
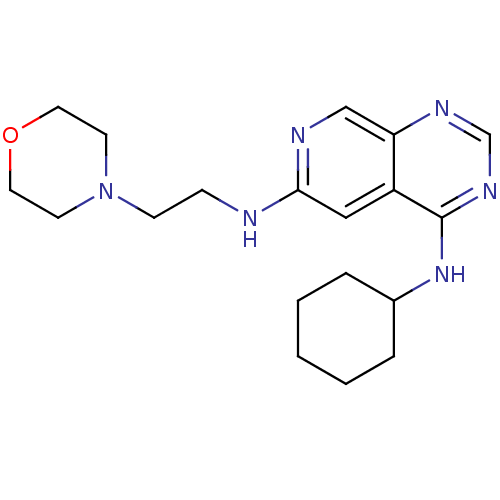
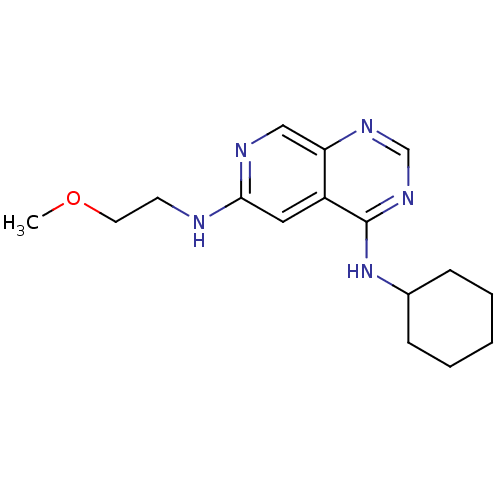
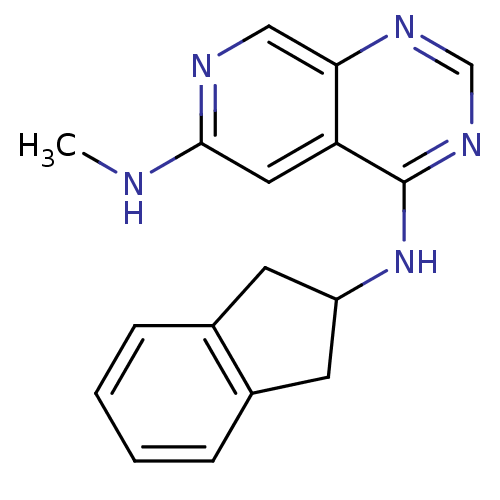
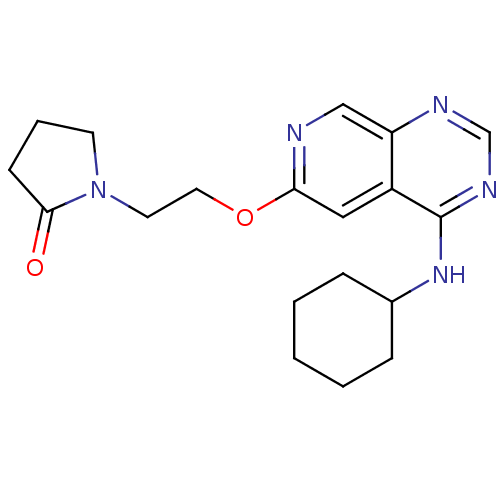
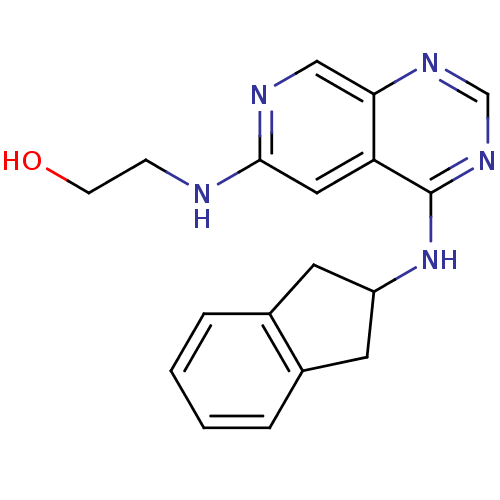
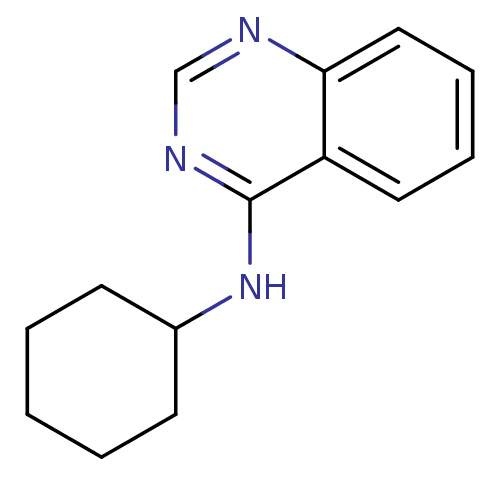
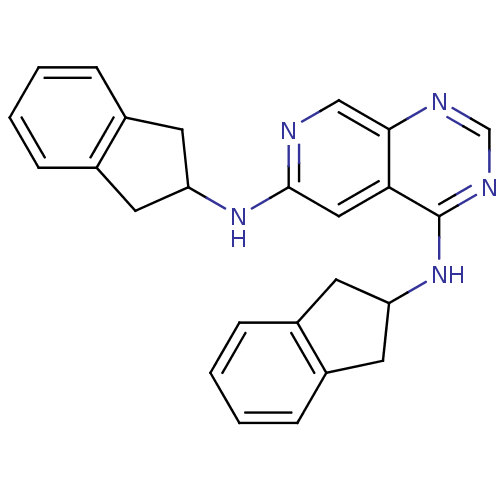
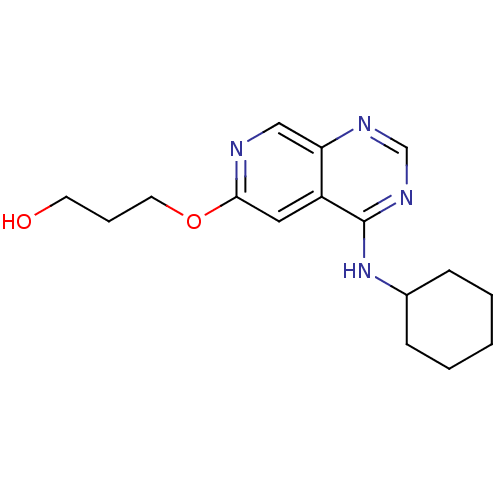
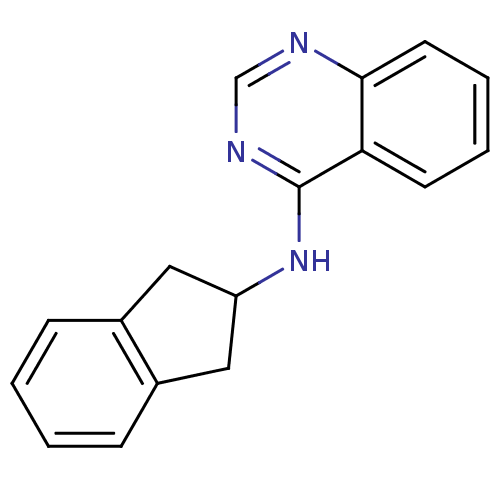
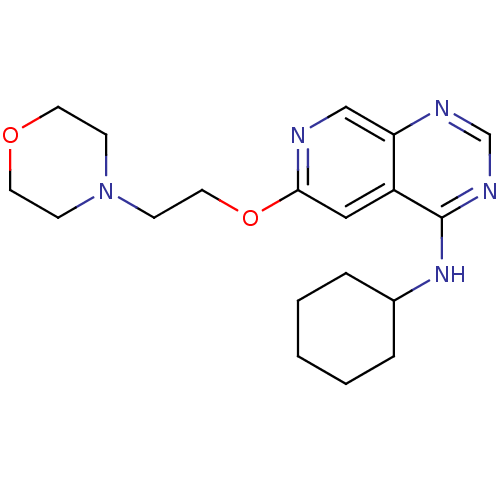
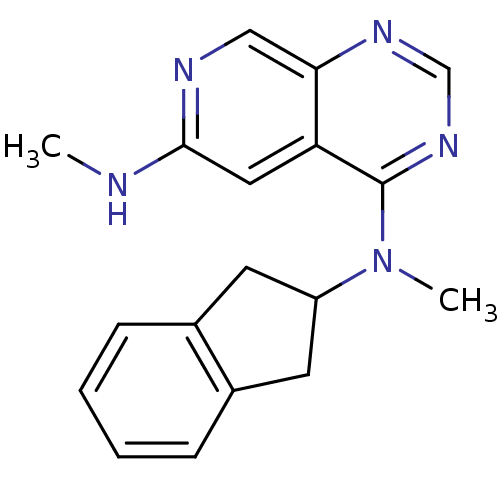
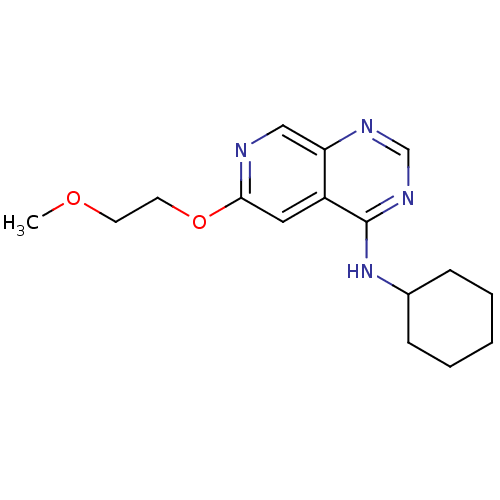
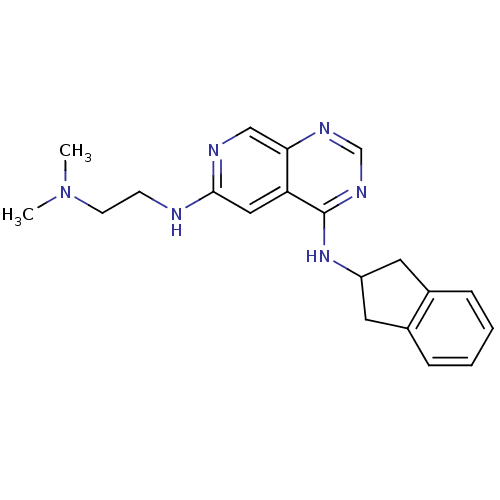
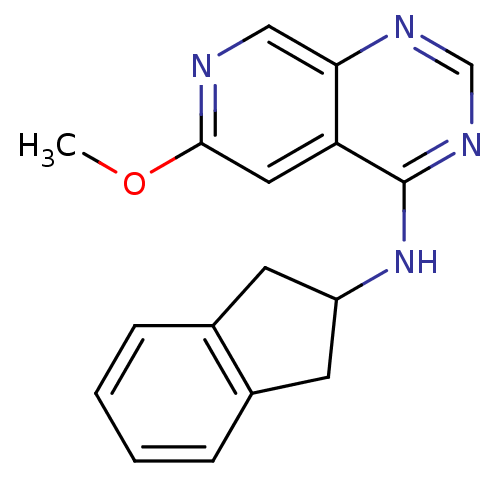
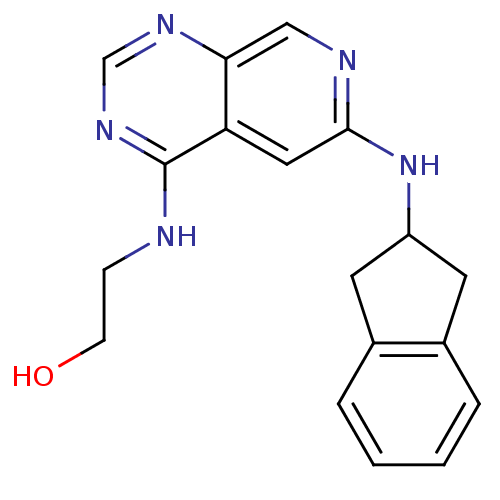
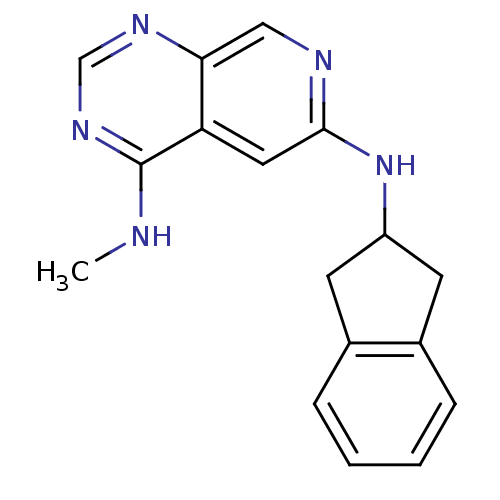
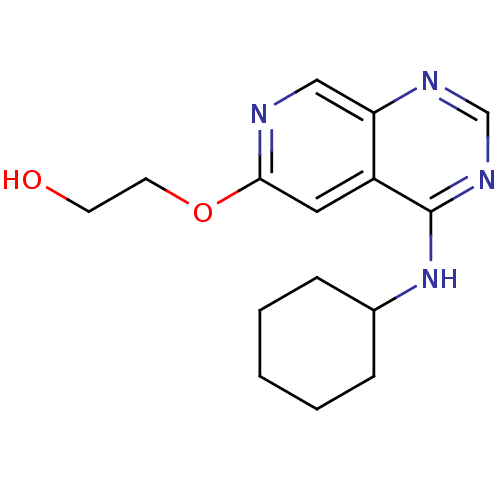
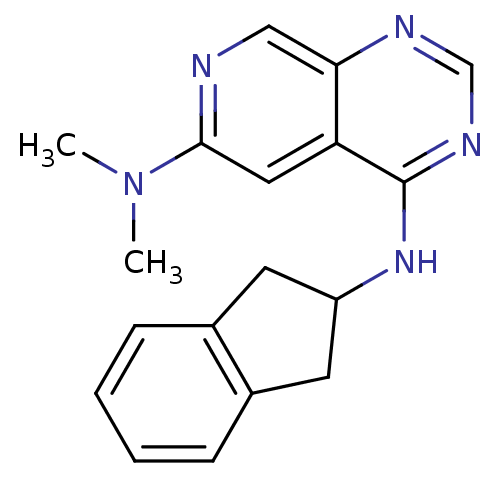
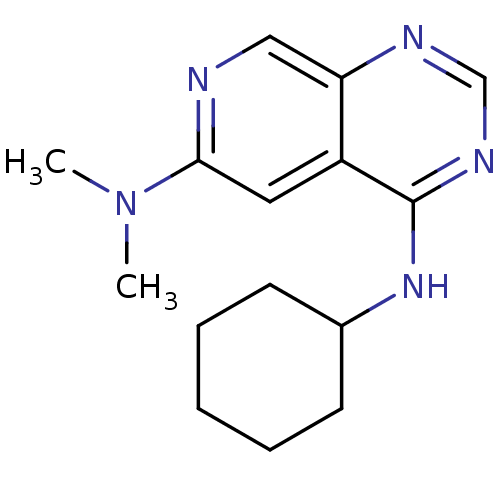
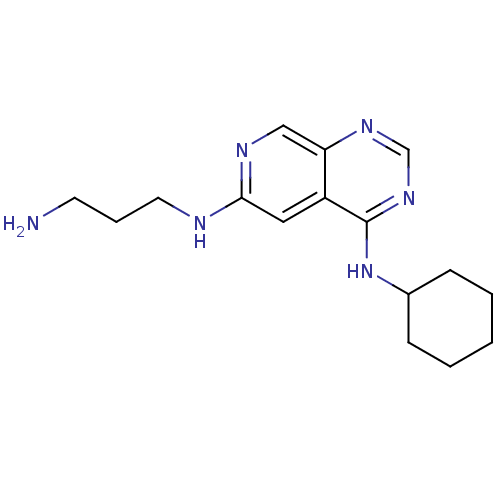
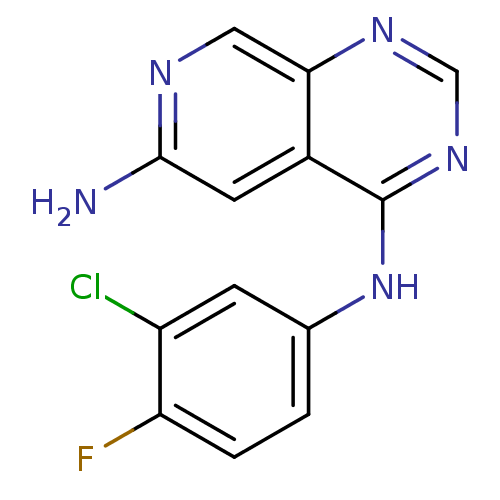
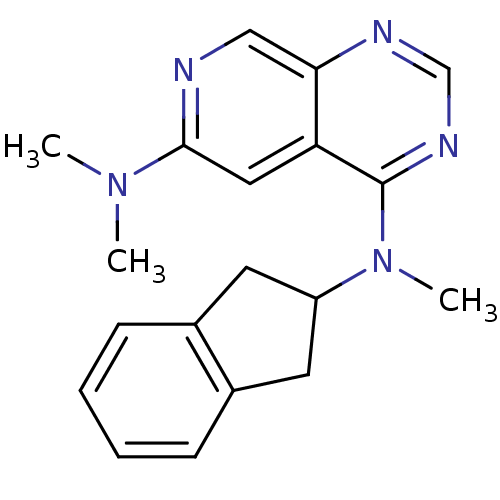
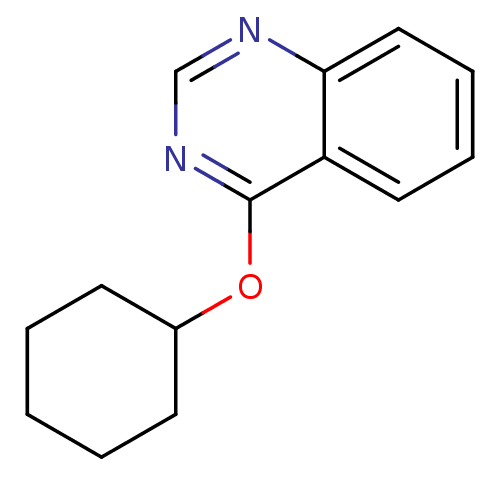
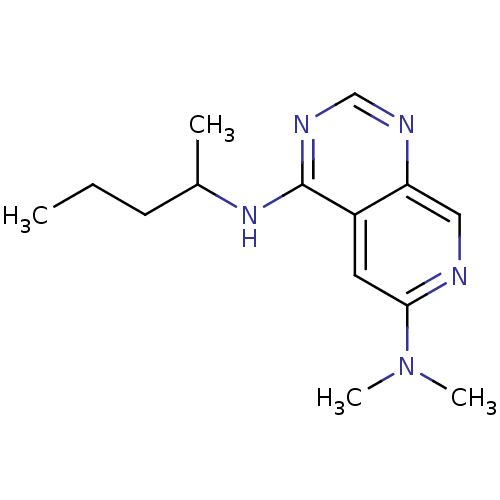
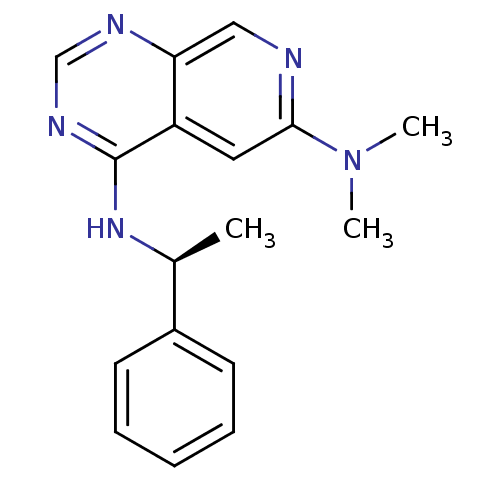
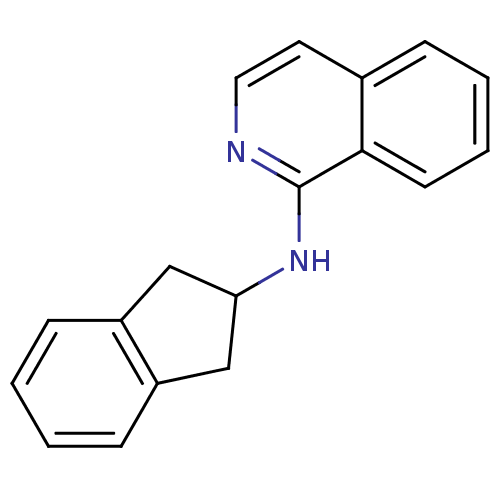
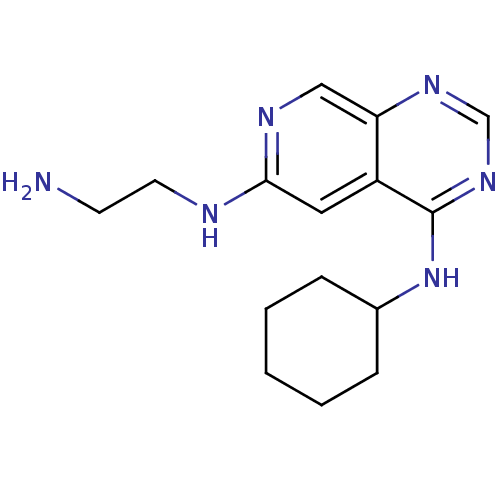
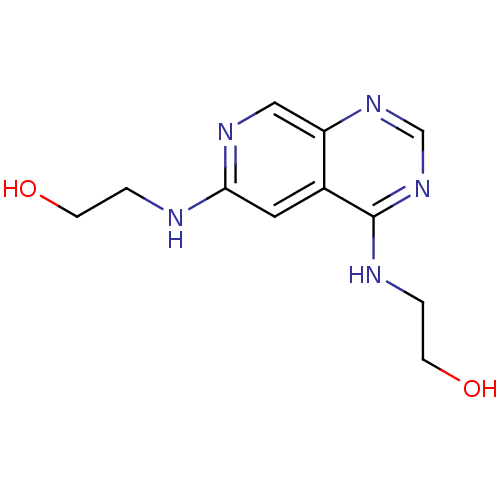
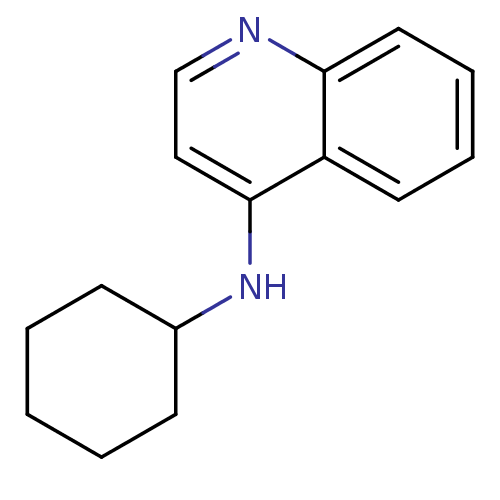
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 50028468
Affinity DataIC50: 3nMAssay Description:Antagonist activity at mGluR1More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 101nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 102nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 115nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 125nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 177nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 185nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 199nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Antagonist activity at mGluR1More data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 694nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 858nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 1.65E+3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Antagonist activity at mGluR5More data for this Ligand-Target Pair
Affinity DataKi: 6.15E+3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair
Affinity DataKi: 6.16E+3nMAssay Description:Antagonist activity at rat mGluR1a expressed in CHO cells assessed as inhibition of glutamate-evoked increase in calcium internalization preincubated...More data for this Ligand-Target Pair