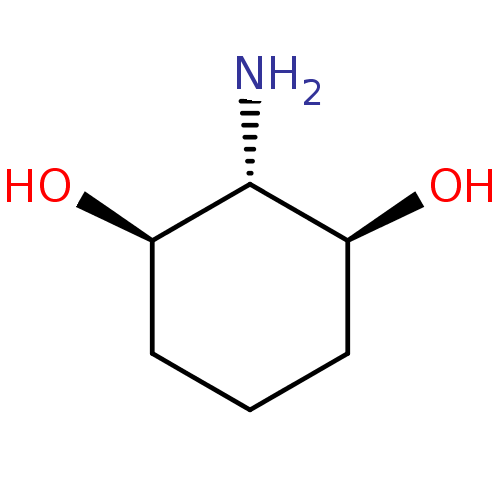
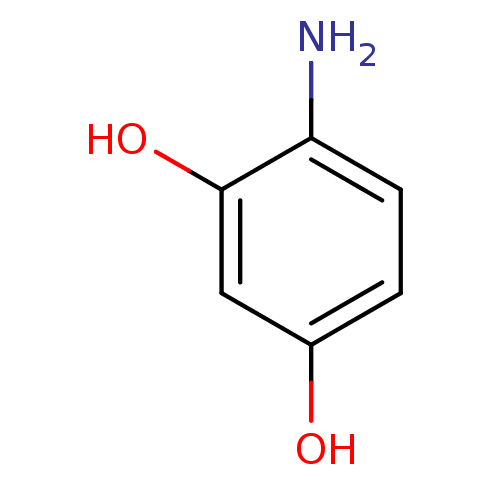
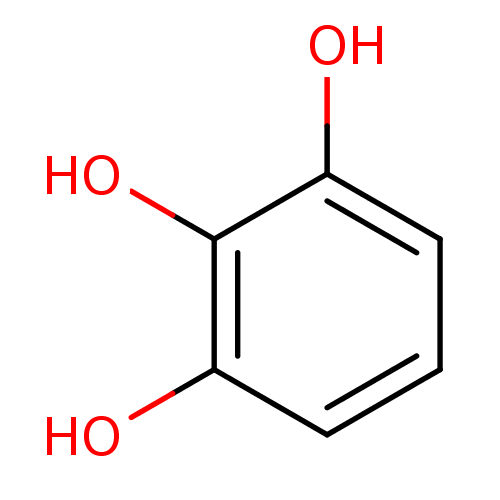
Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 50038271
Affinity DataIC50: 4.40nMAssay Description:Inhibition of rat intestinal maltaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of rat intestinal maltaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMAssay Description:Inhibition of rat intestinal maltaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nMAssay Description:Inhibition of sucrose hydrolyzing activity of rat intestinal alpha glucosidaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+4nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of rat intestinal sucraseMore data for this Ligand-Target Pair