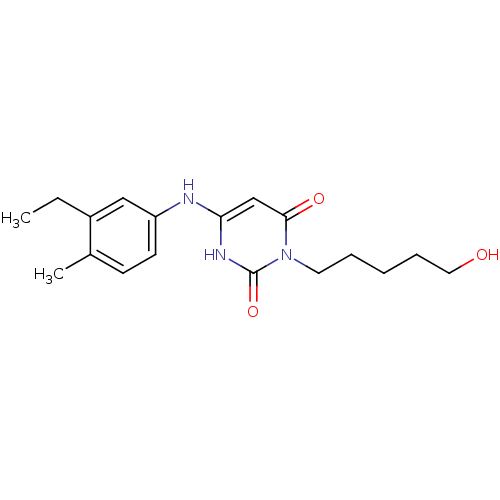
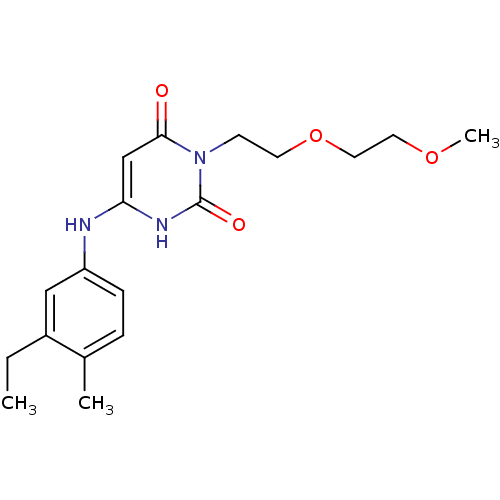
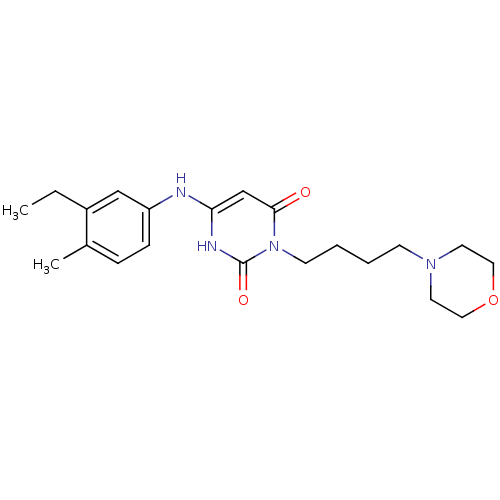
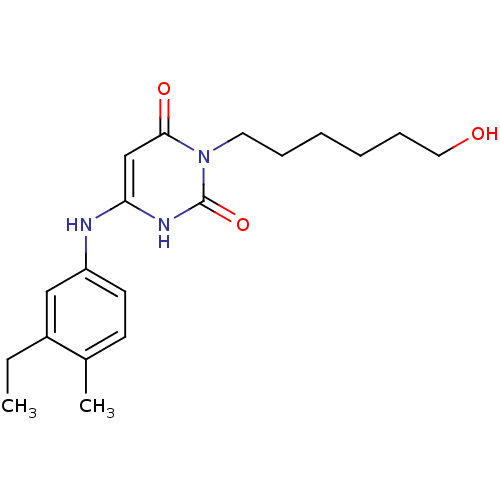
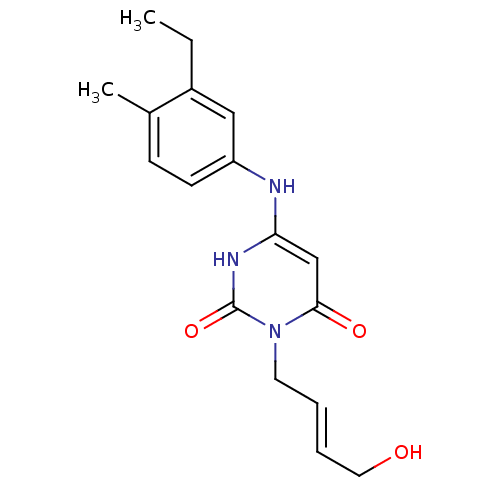
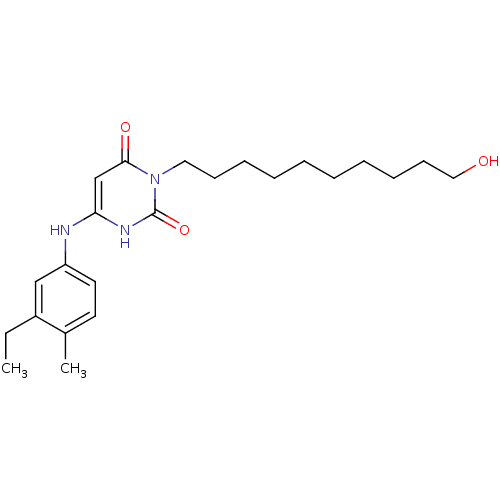
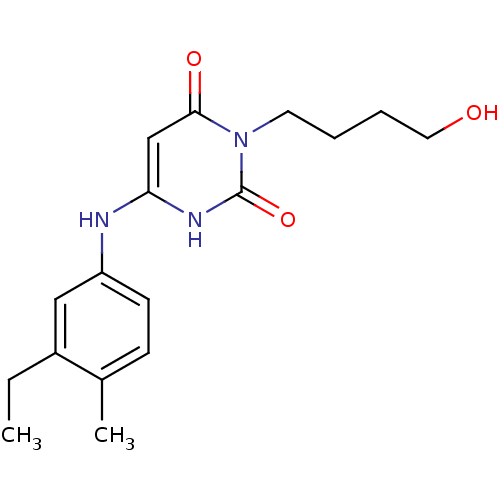
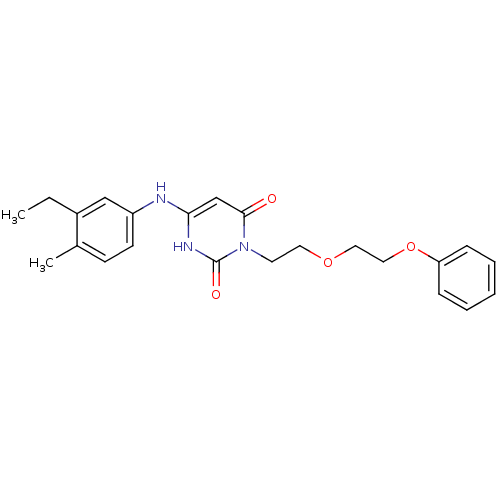
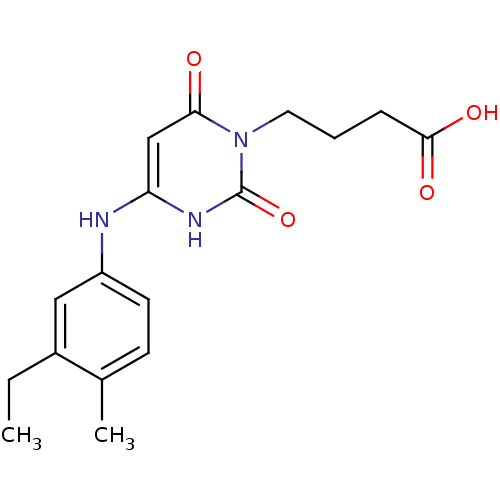
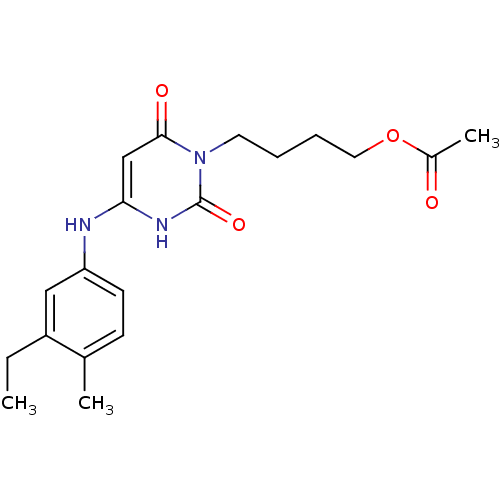
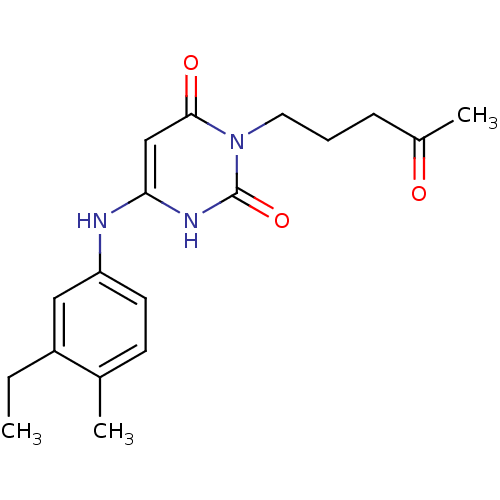
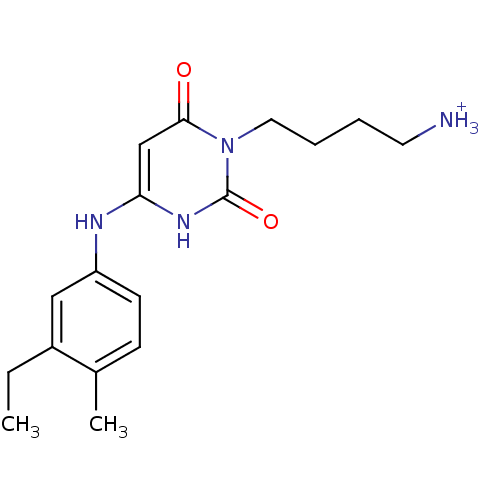
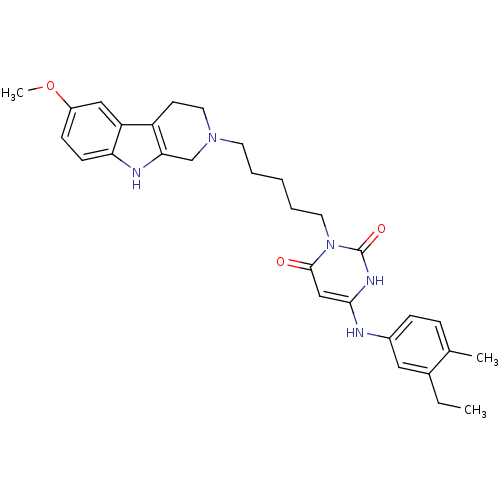
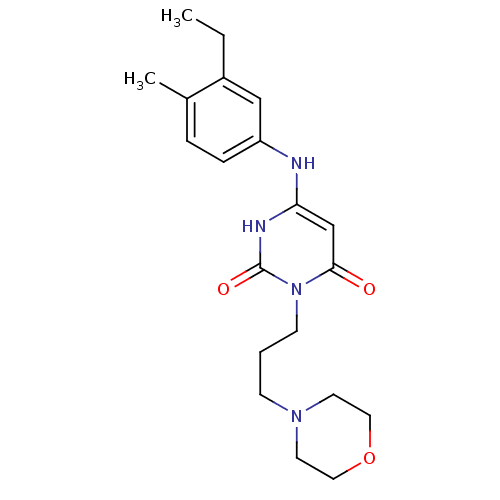
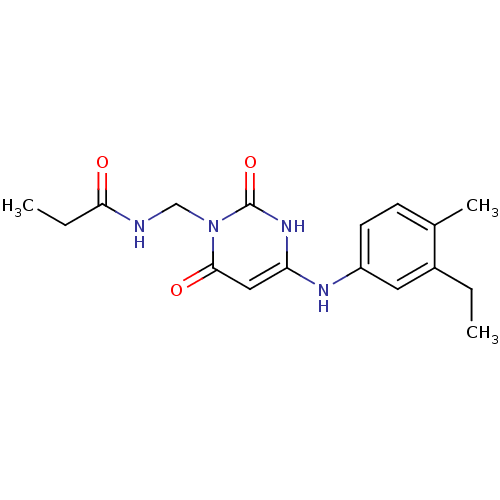
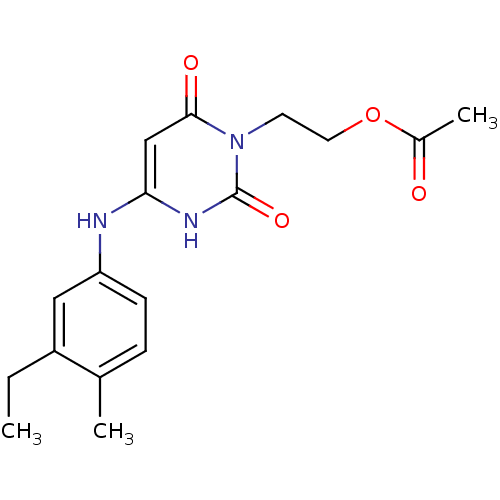
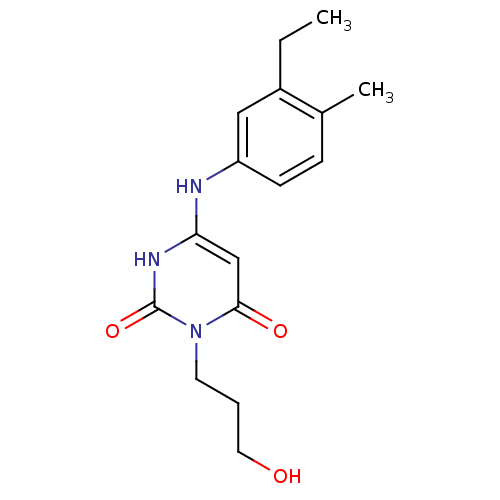
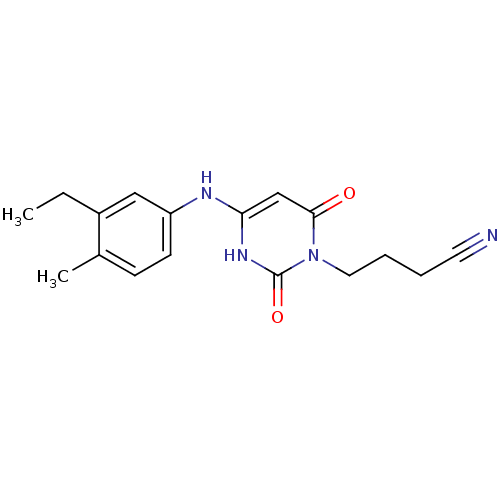
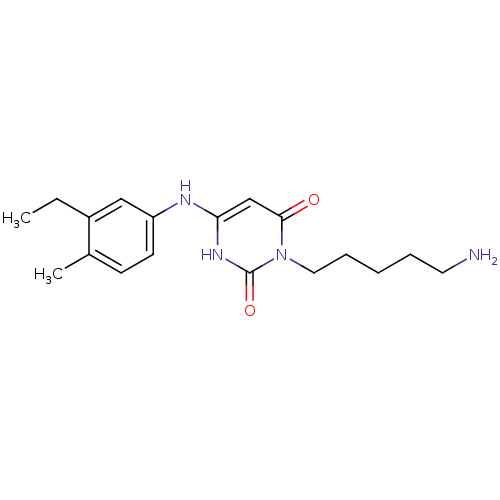
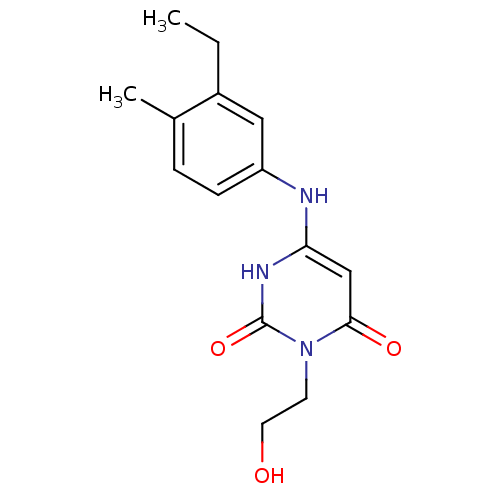
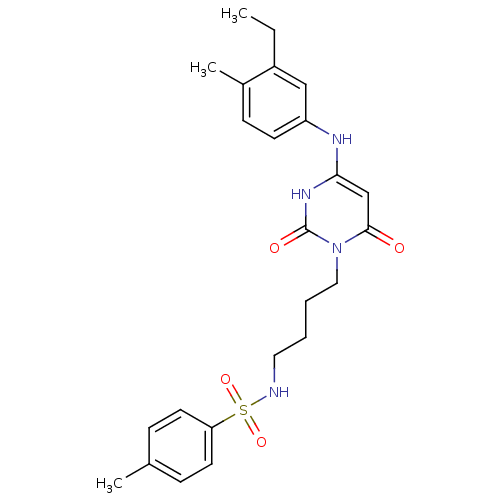
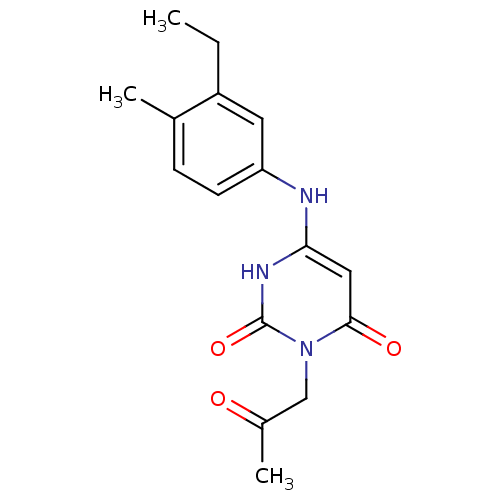
Report error Found 47 Enz. Inhib. hit(s) with all data for entry = 50037628
Affinity DataKi: 21nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 49nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 58nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 71nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 75nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 82nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 87nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 87nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 88nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 96nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 98nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 125nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 185nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 230nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Inhibitory concentration against Bacillus subtilis DNA polymerase IIIC using [3H]dTMP 250 pM (30 degree C for 10 min)More data for this Ligand-Target Pair