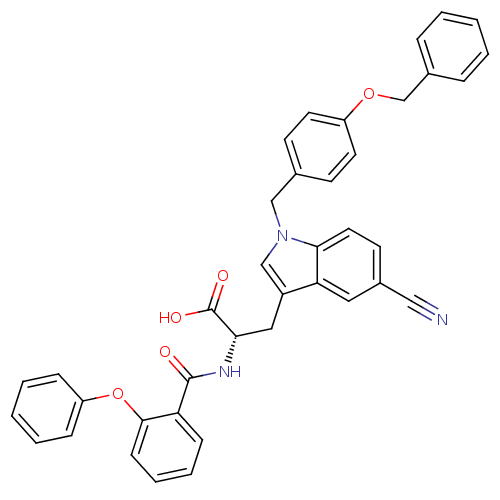
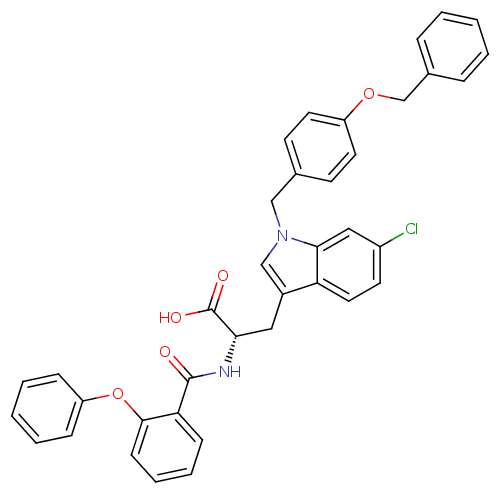
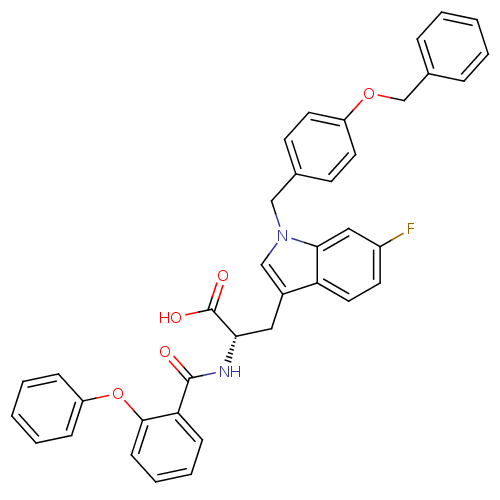
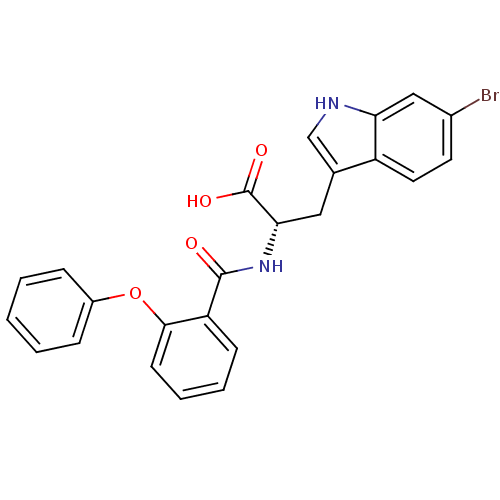
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 3311
Affinity DataKi: 100nM ΔG°: -39.7kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -38.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 400nM ΔG°: -36.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -35.7kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 600nM ΔG°: -35.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 800nM ΔG°: -34.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM ΔG°: -34.0kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nM ΔG°: -31.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nM ΔG°: -29.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -26.6kJ/molepH: 7.4 T: 2°CAssay Description:Ki values were determined from a competition experiment in which serial dilutions of inhibitor were added to compete against a fixed concentration (1...More data for this Ligand-Target Pair