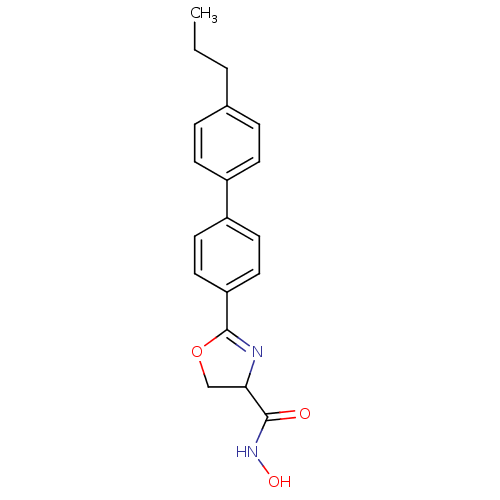
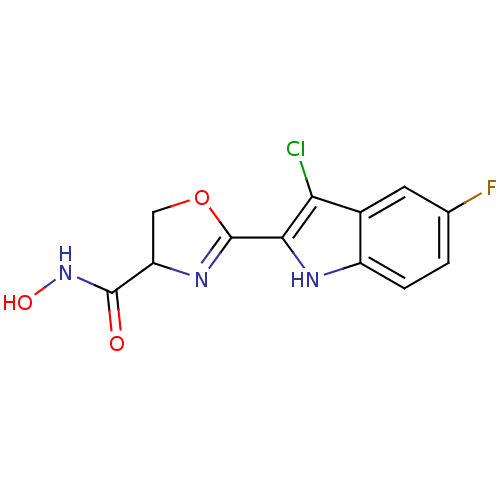
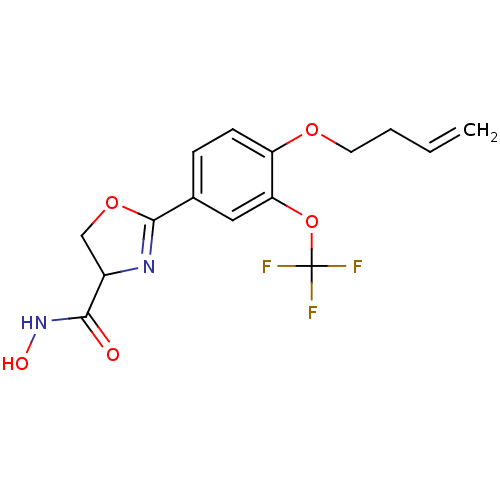
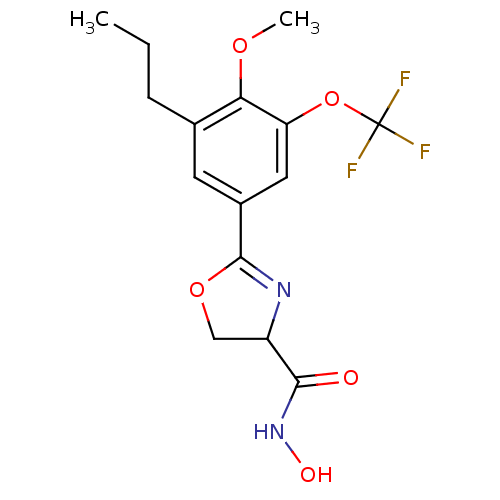
Report error Found 64 Enz. Inhib. hit(s) with all data for entry = 50012151
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
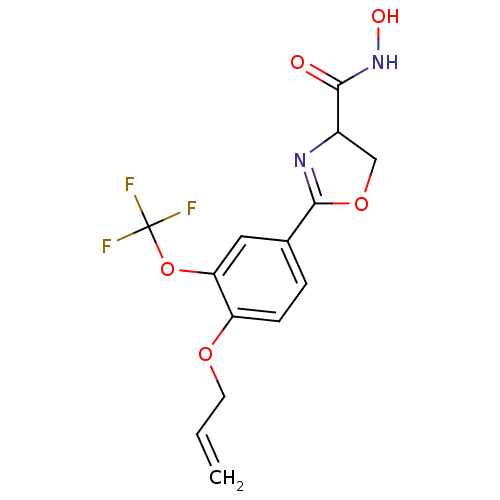
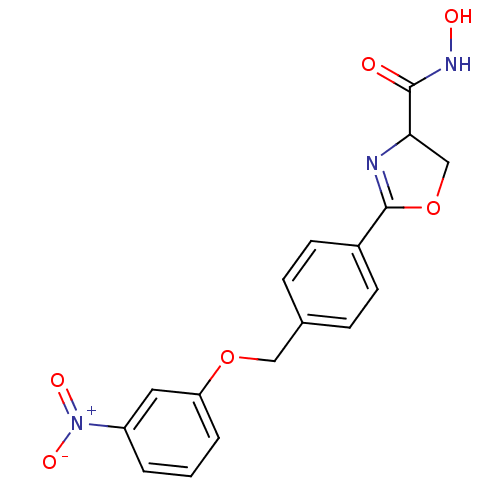
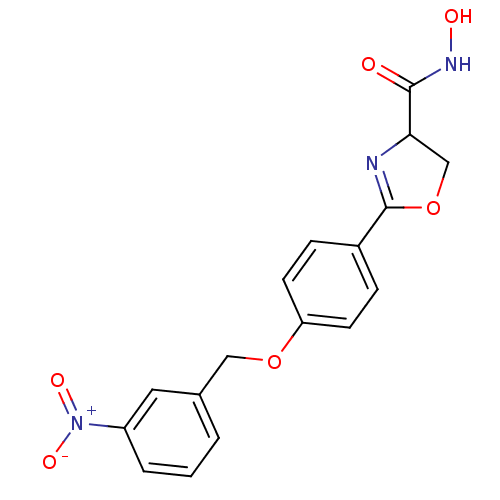
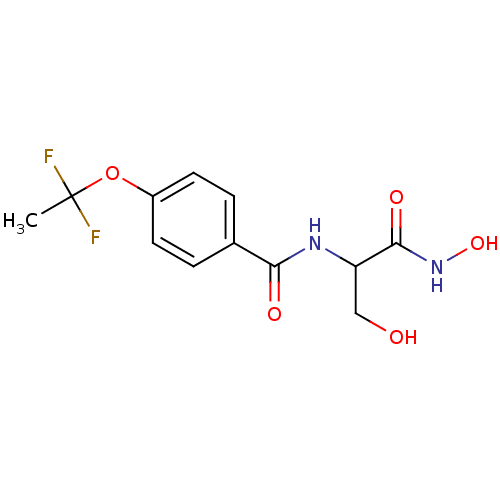
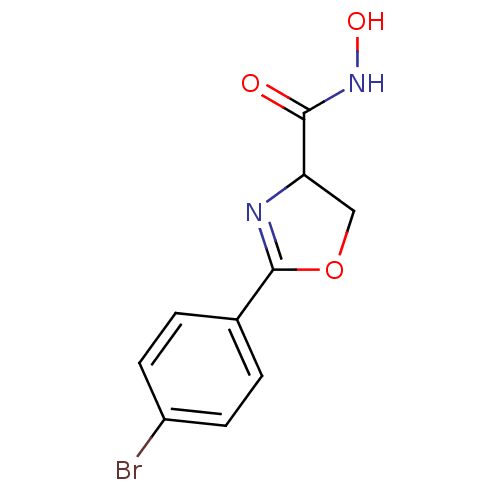
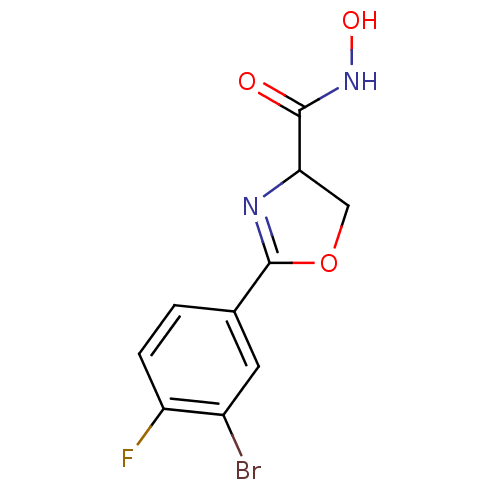
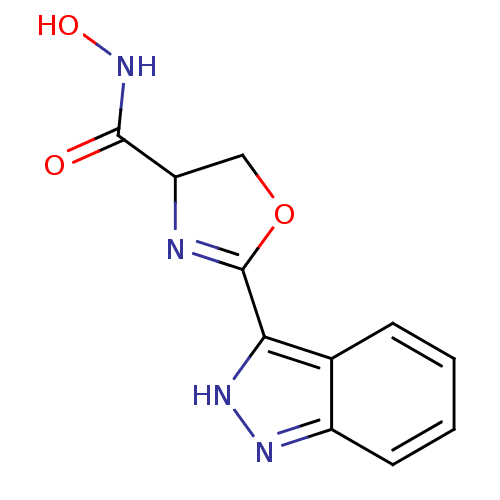
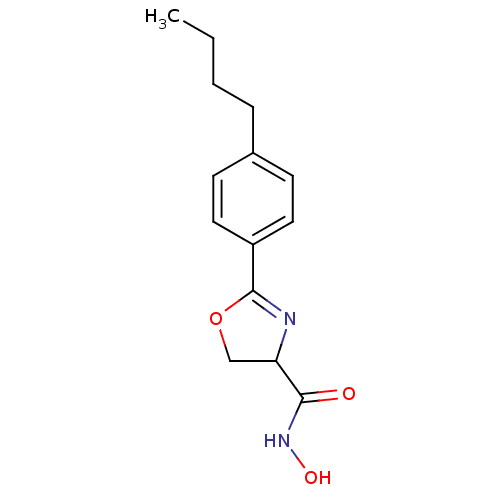
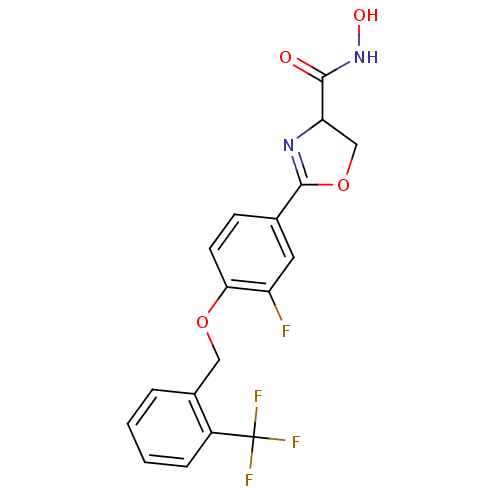
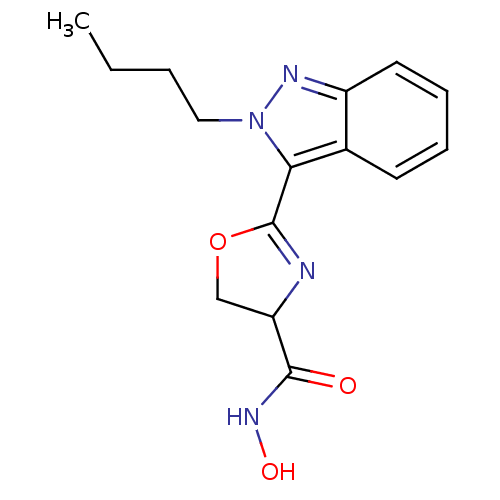
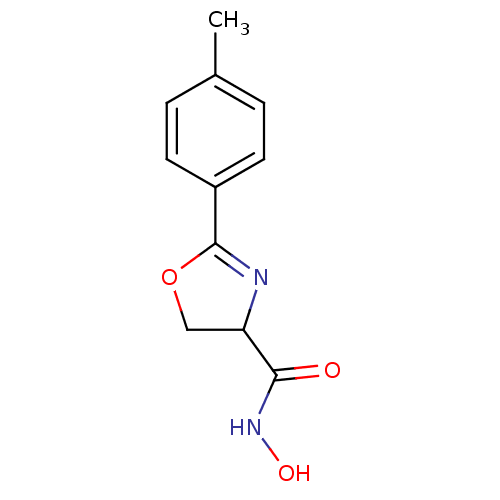
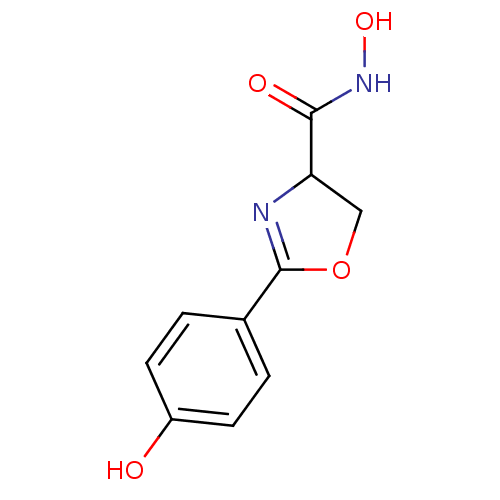
Affinity DataIC50: 120nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
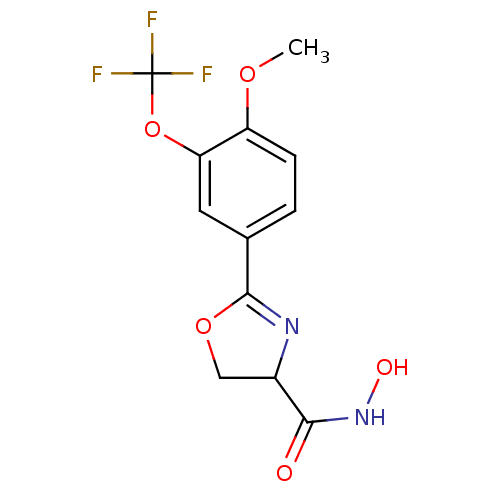
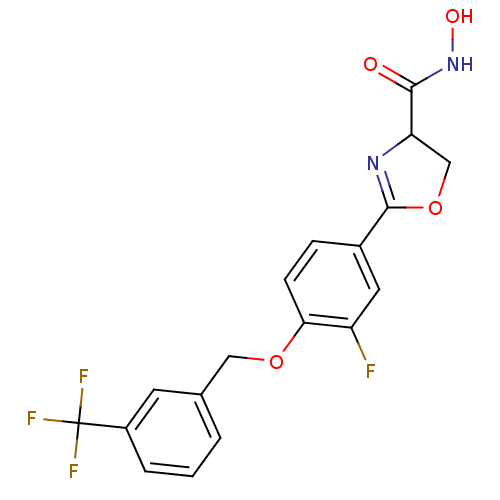
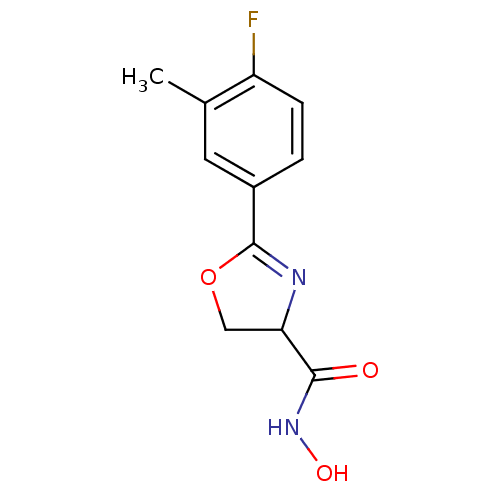
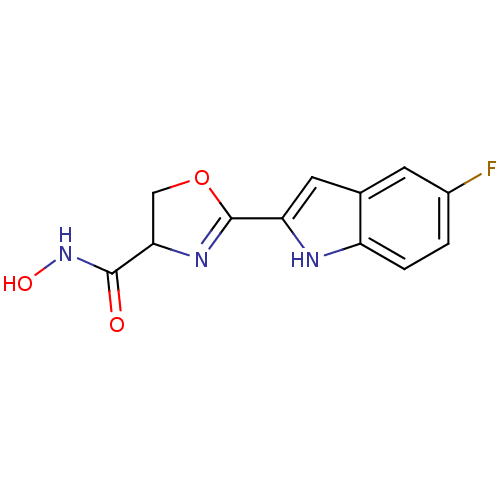
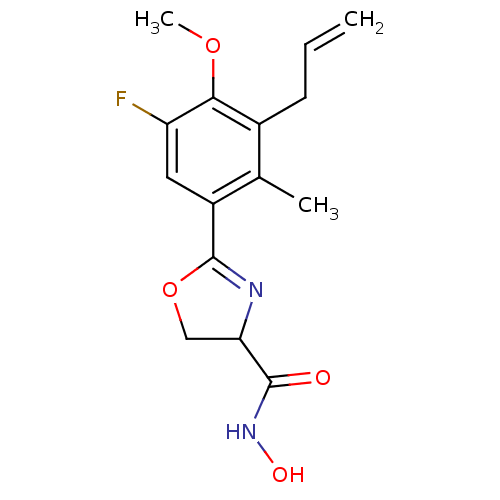
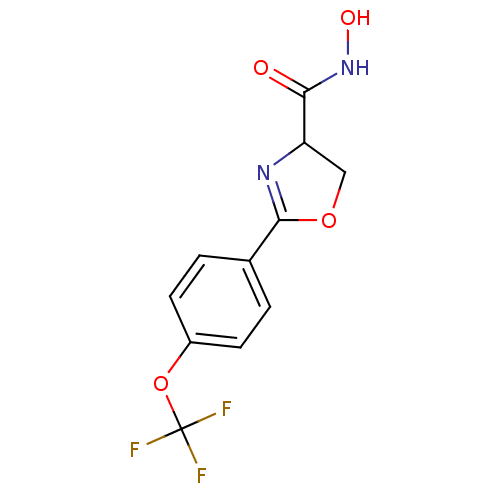
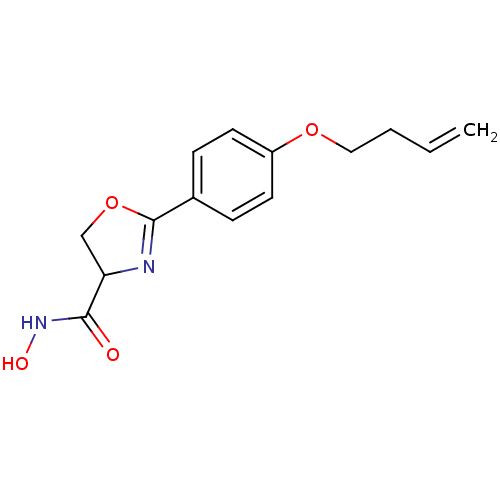
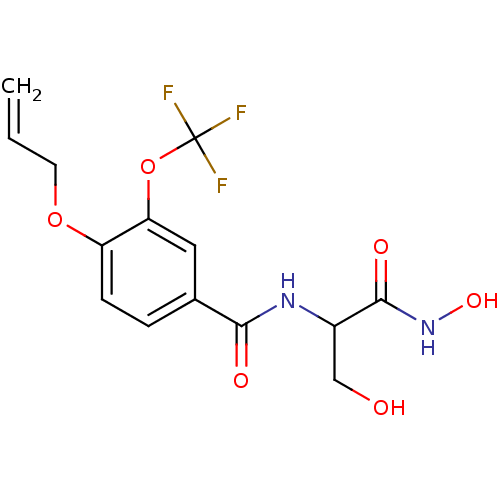
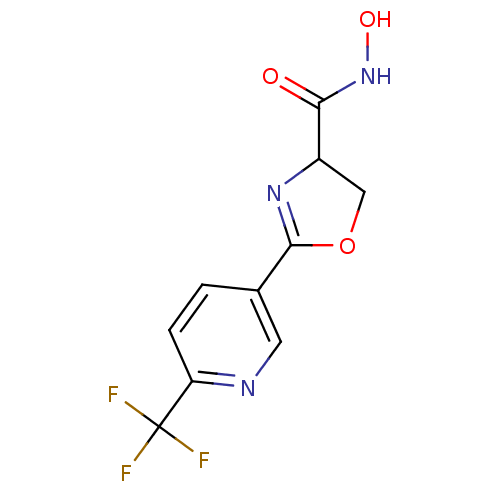
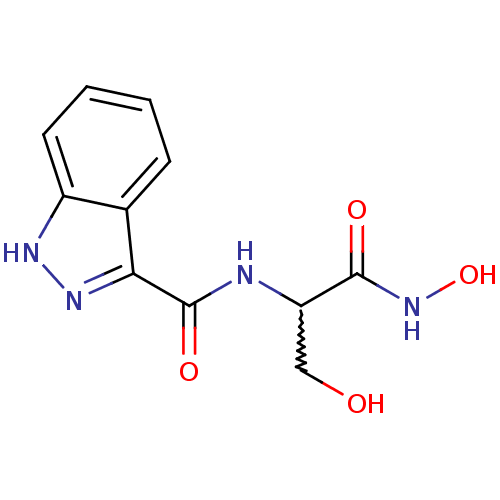
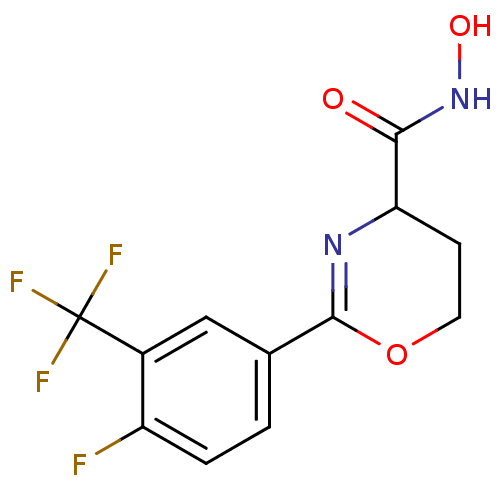
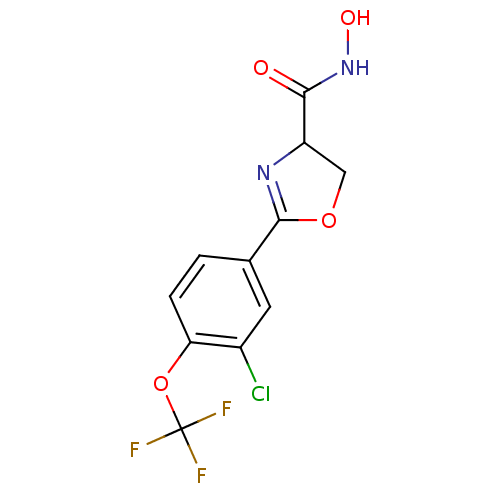
Affinity DataIC50: 160nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
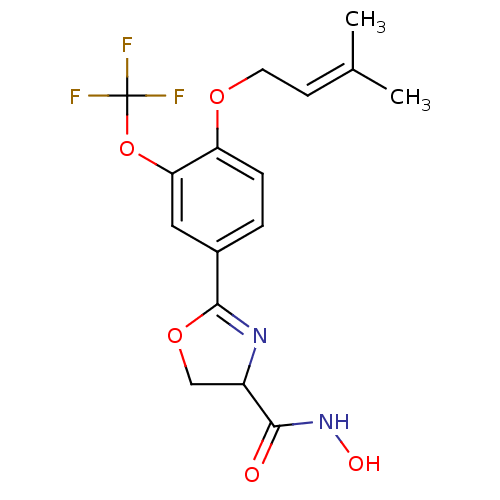
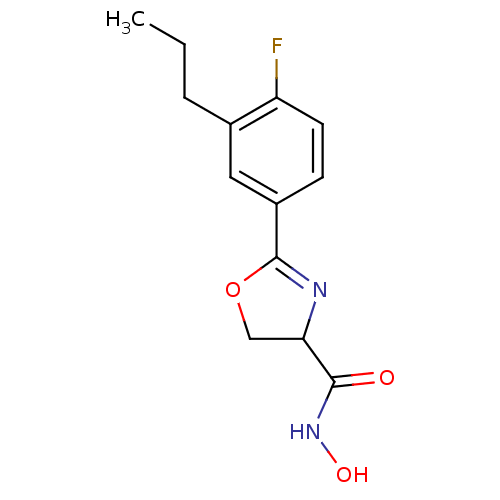
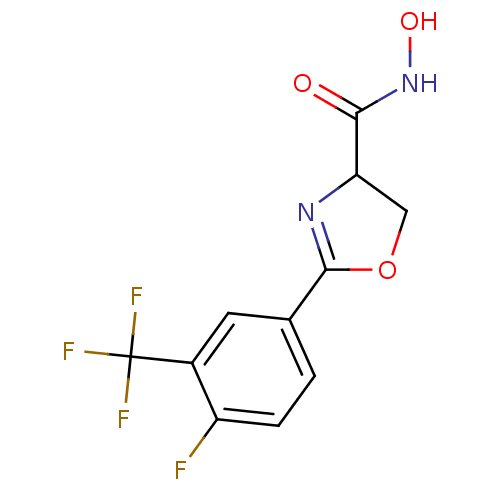
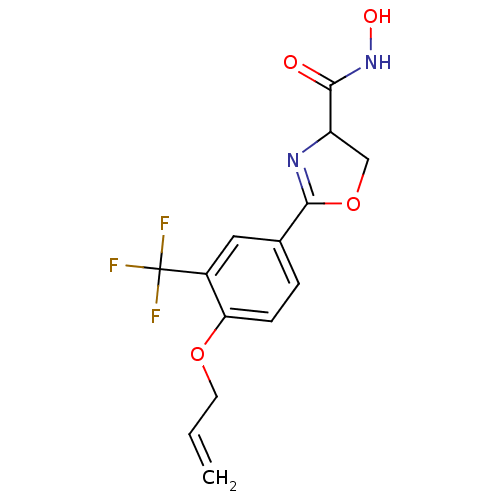
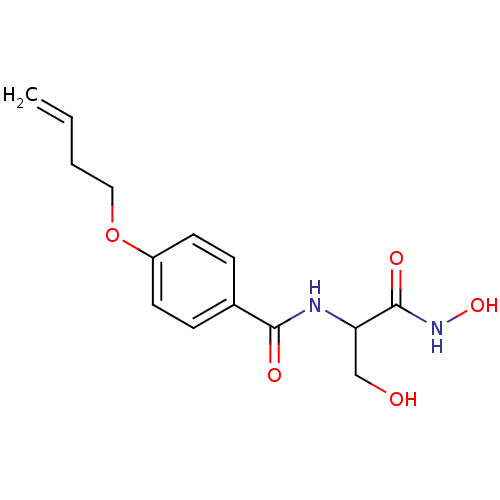
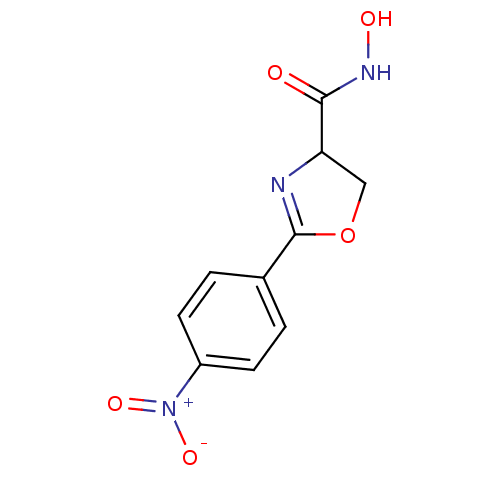
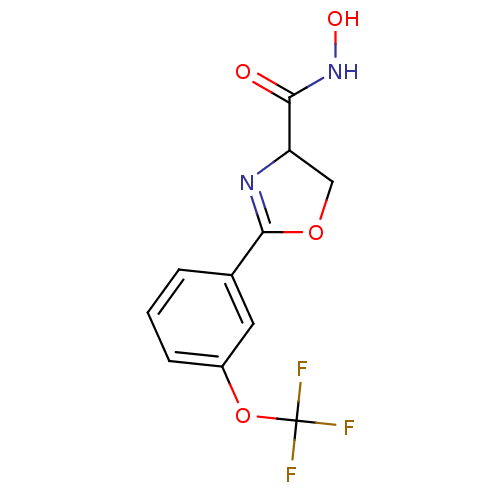
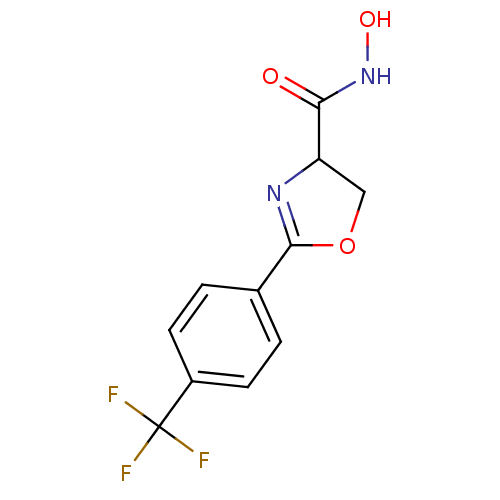
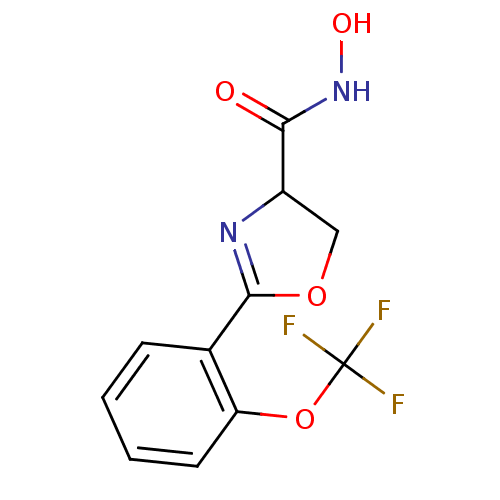
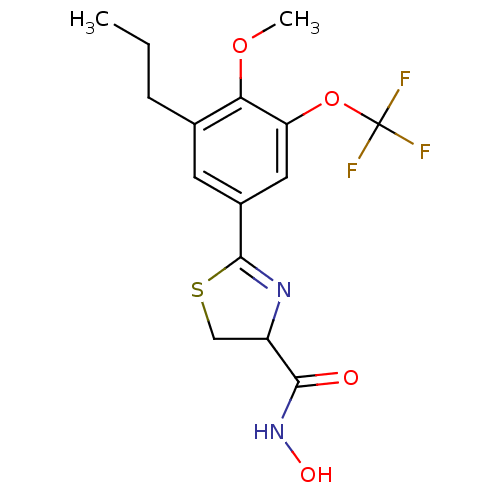
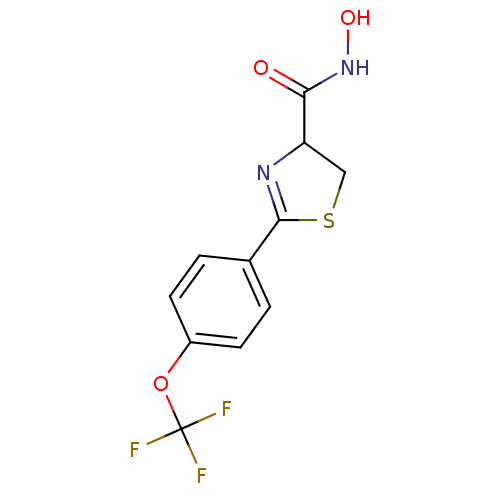
Affinity DataIC50: 190nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
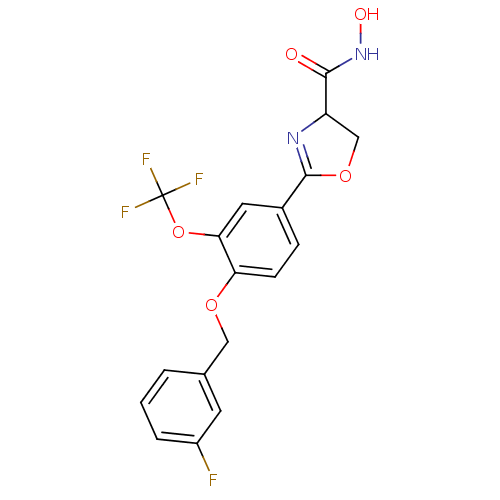
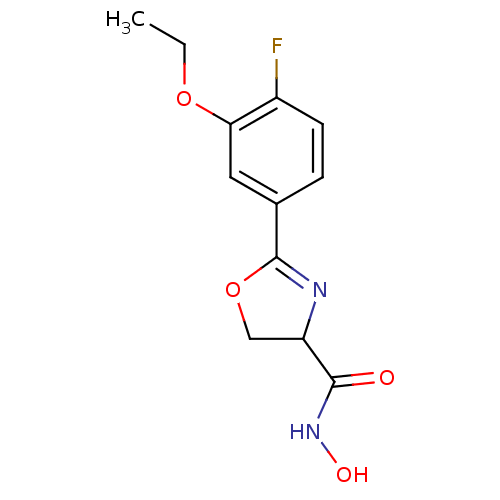
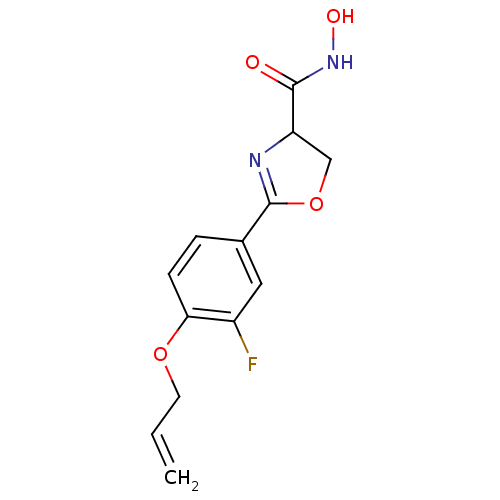
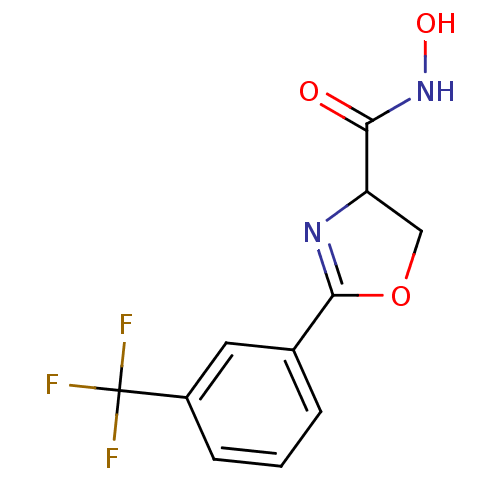
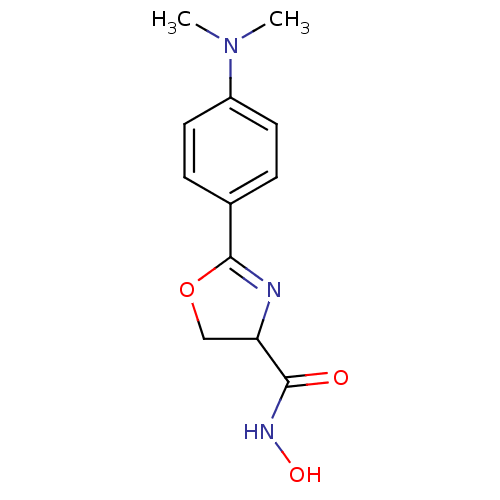
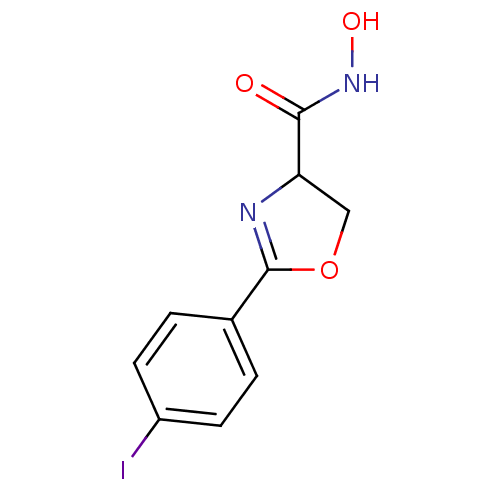
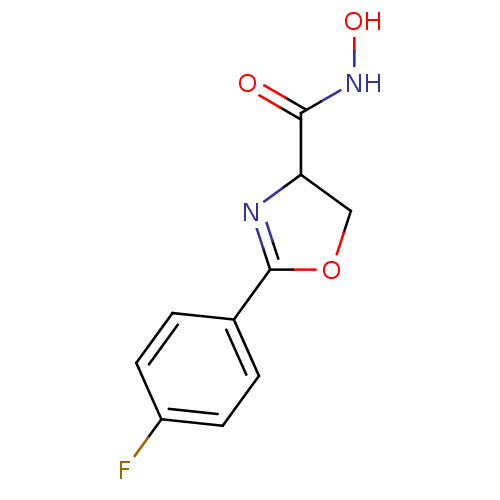
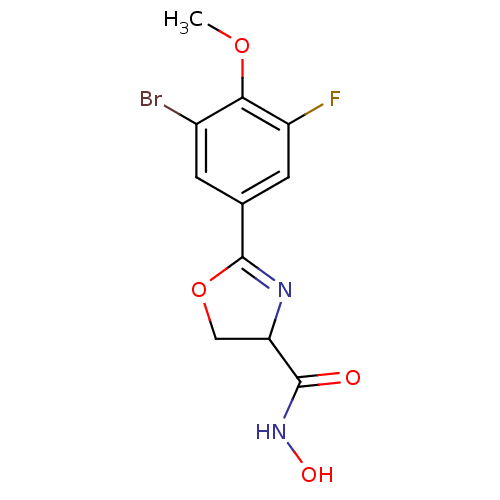
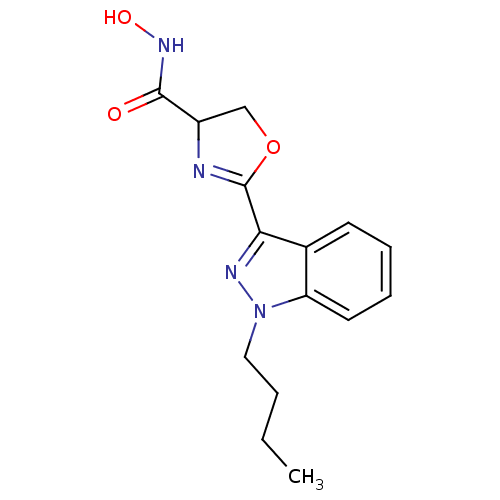
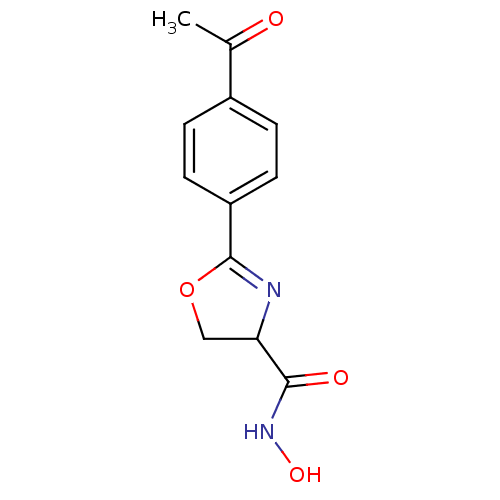
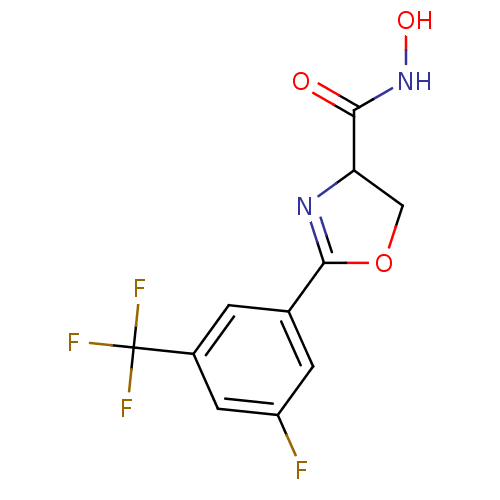
Affinity DataIC50: 250nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 350nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 560nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 840nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 960nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.54E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 3.90E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 5.60E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 7.50E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 7.80E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 9.60E+3nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.78E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.12E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Chiron
Curated by ChEMBL
Chiron
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Pseudomonas aeruginosa UDP-3-O-acyl-N-acetylglucosamine Deacetylase (LpxC) in vitro.More data for this Ligand-Target Pair