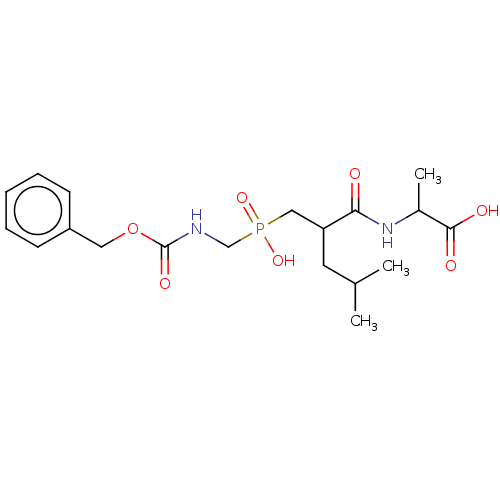
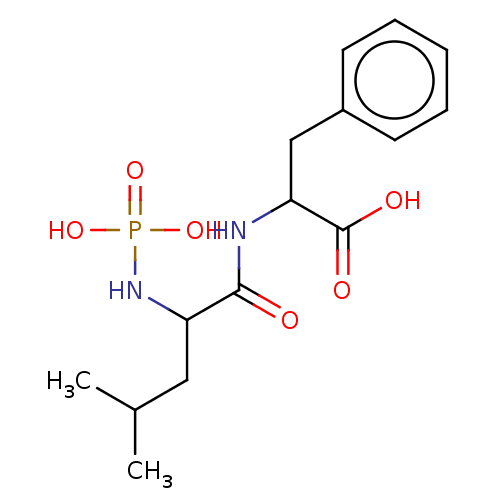
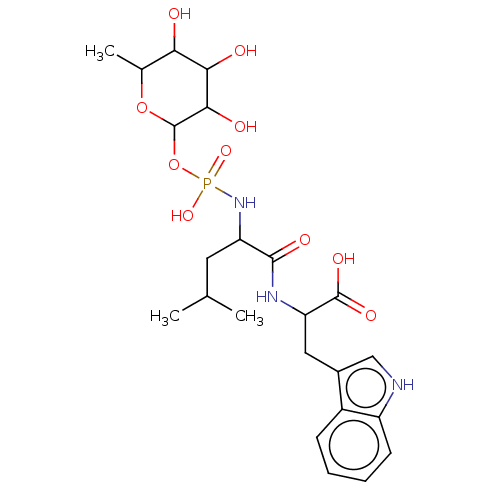
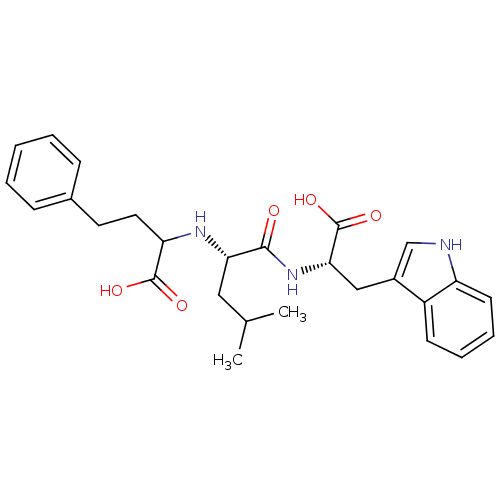
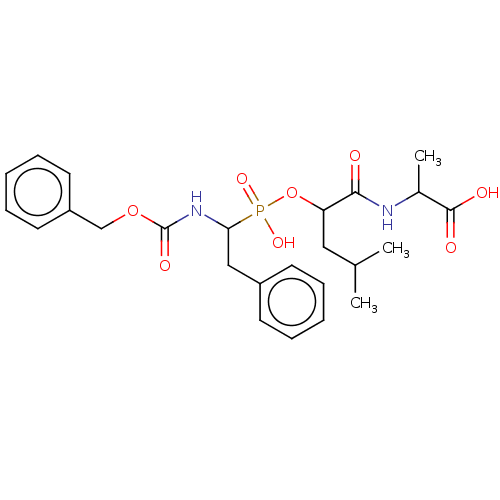
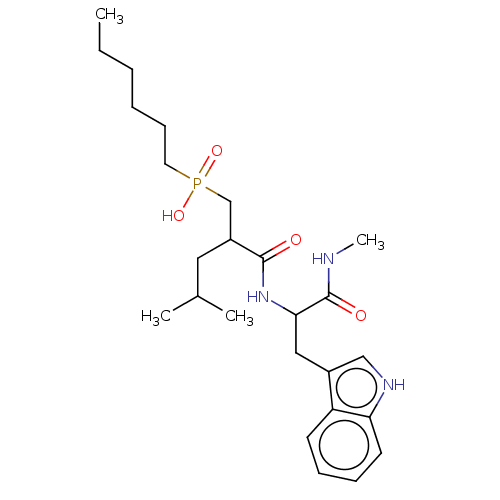
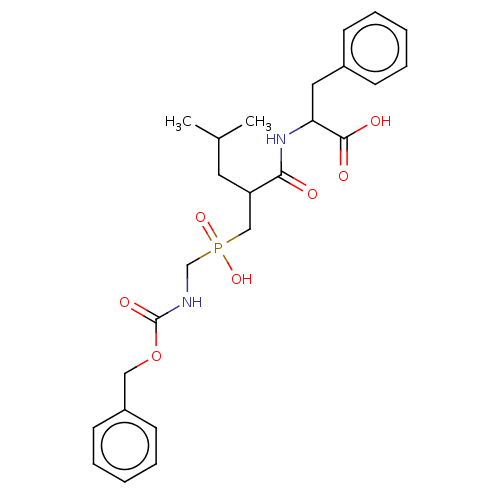
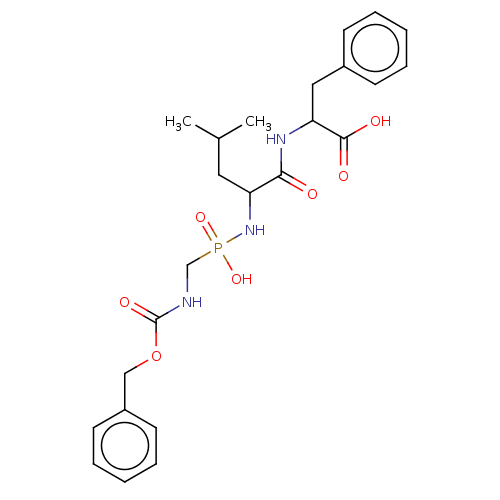
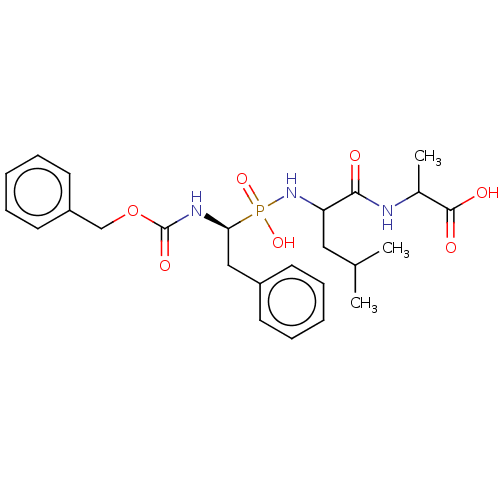
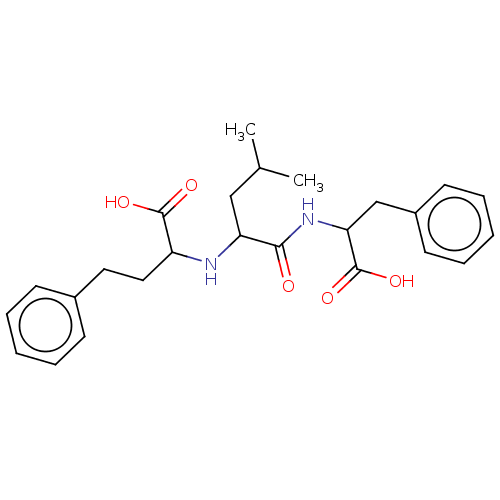
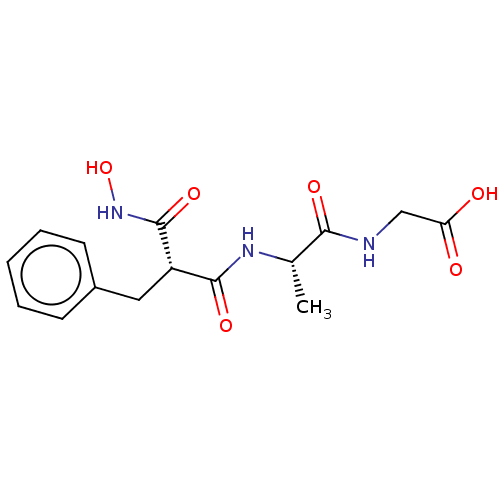
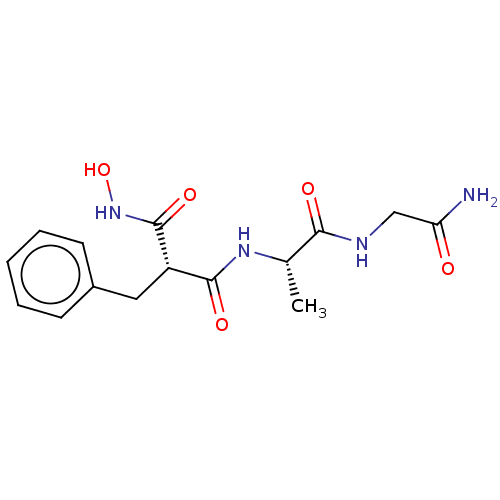
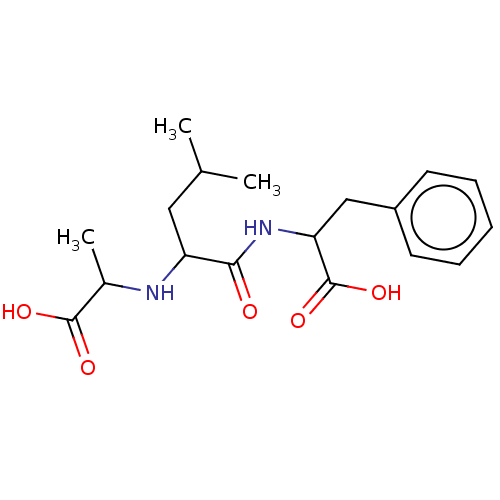
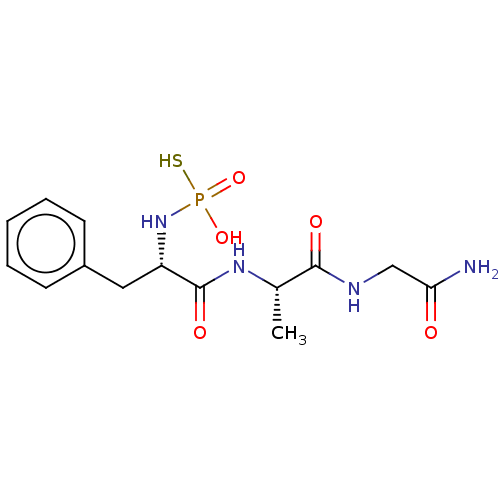
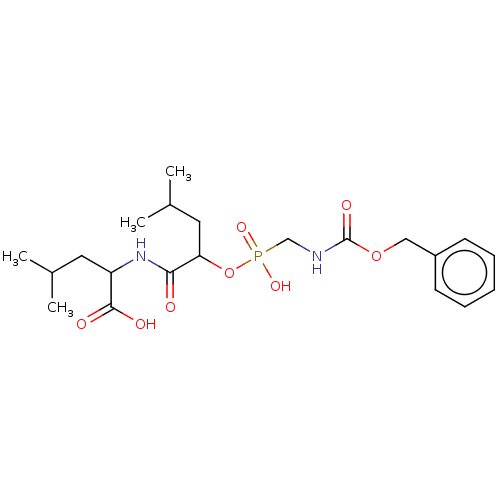
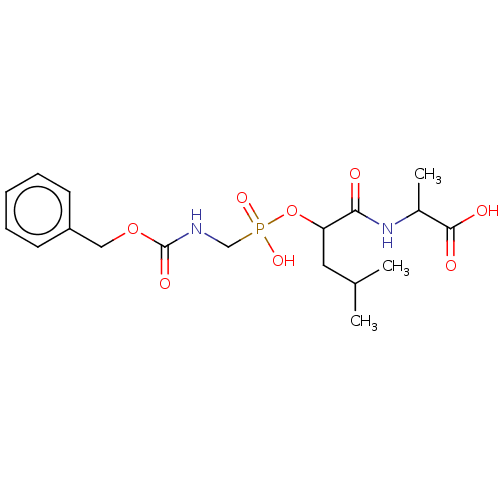
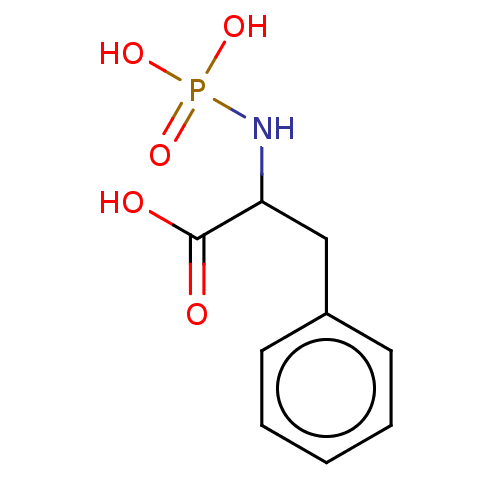
Report error Found 76 Enz. Inhib. hit(s) with all data for entry = 50003633
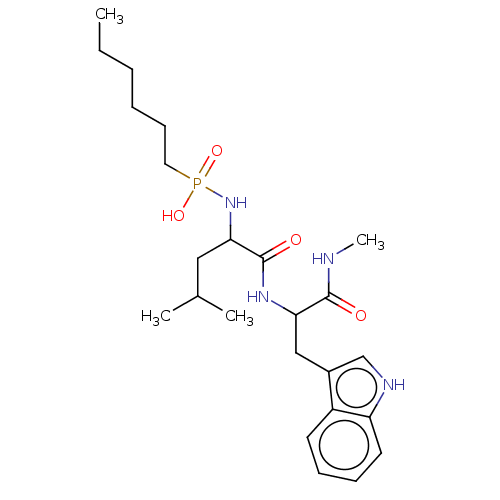
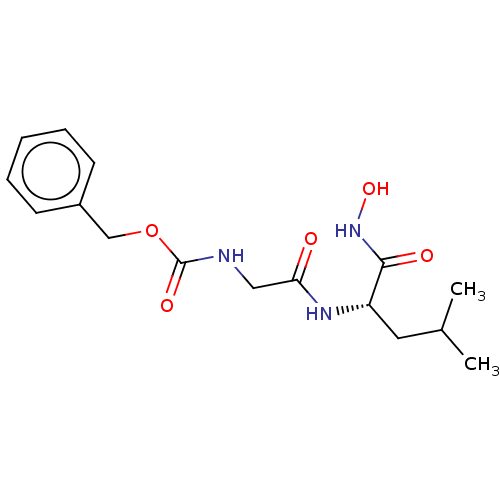
Affinity DataKi: 0.0680nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
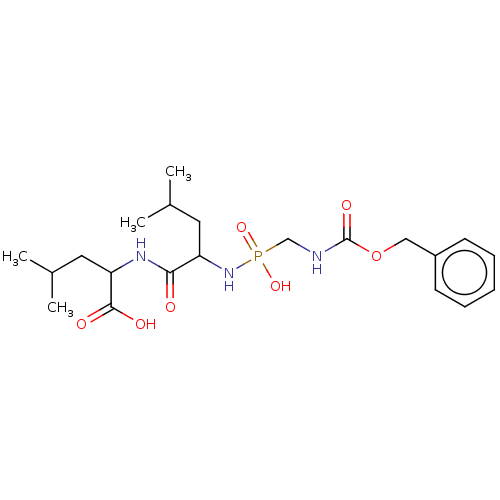
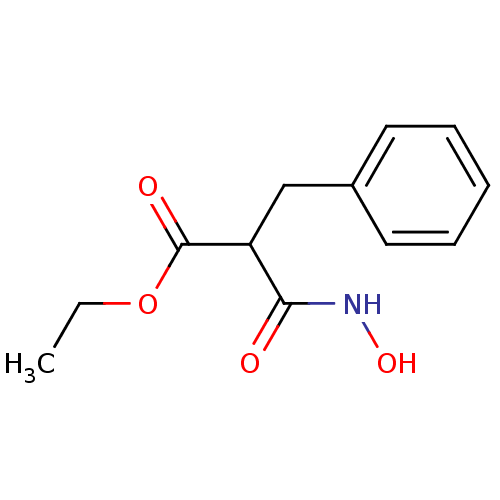
Affinity DataKi: 1.51nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
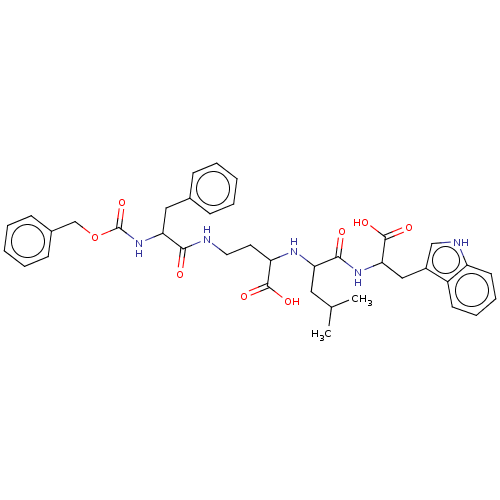
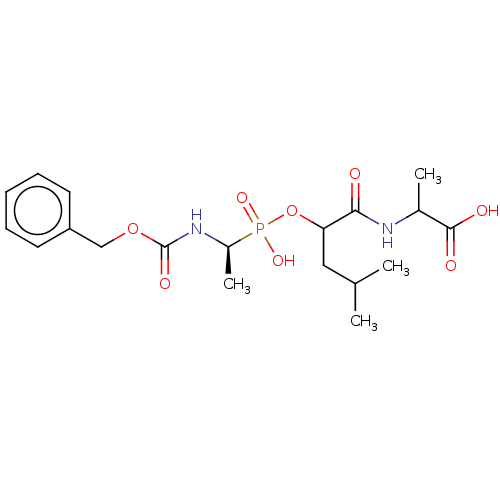
Affinity DataKi: 9.12nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
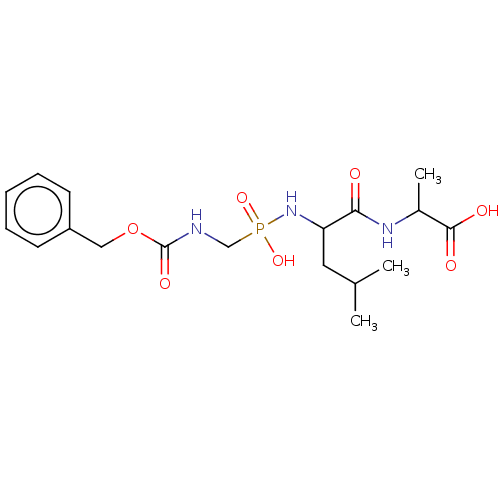
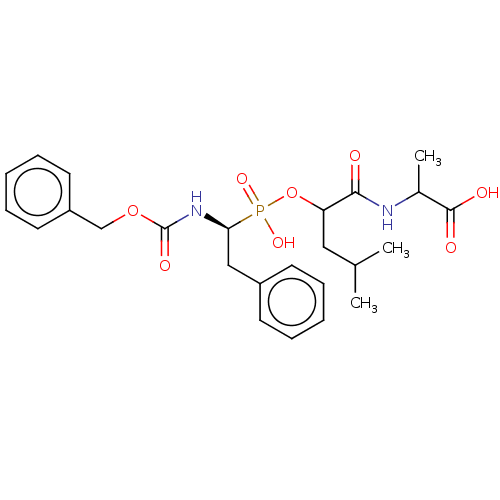
Affinity DataKi: 11nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 16.6nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 18.6nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 19.1nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 28.2nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 33.9nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 44.7nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 52.5nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 66.1nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 75.9nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 181nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 218nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 269nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 301nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 363nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 407nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 426nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 478nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 478nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 602nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 660nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 660nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 676nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 758nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 758nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 851nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.41E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.44E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.69E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.81E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.81E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 2.29E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 2.57E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 2.69E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 2.81E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 6.91E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 8.91E+3nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.28E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.28E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 1.99E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 2.39E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 3.01E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 3.89E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 4.16E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 4.78E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 5.37E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
Affinity DataKi: 7.24E+4nMAssay Description:pKi value expresses binding affinity against thermolysin pKi = -logKi (where Ki is in mol/L)More data for this Ligand-Target Pair
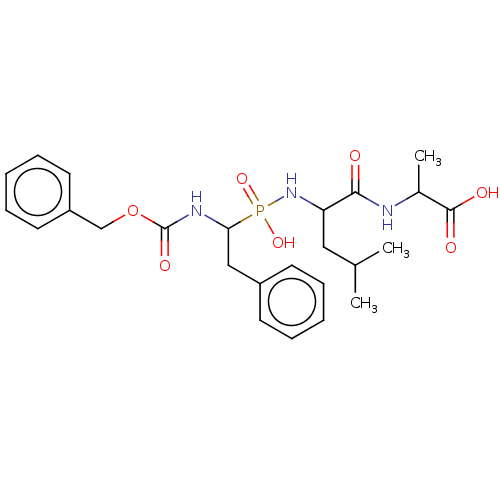
 3D Structure (crystal)
3D Structure (crystal)