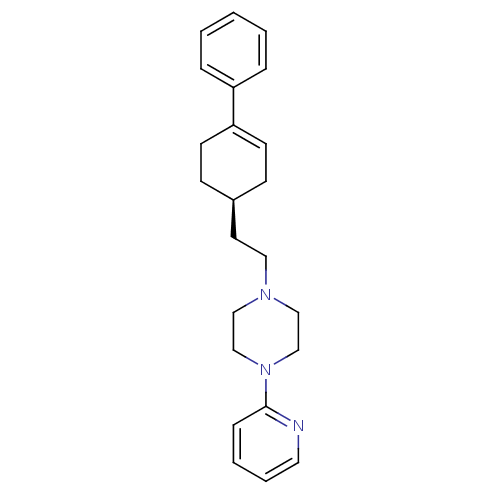
BDBM50055724 1-[2-((S)-4-Phenyl-cyclohex-3-enyl)-ethyl]-4-pyridin-2-yl-piperazine::CHEMBL418937::PD-137789
SMILES C(CN1CCN(CC1)c1ccccn1)[C@H]1CCC(=CC1)c1ccccc1
InChI Key InChIKey=MQHGACGPMGUHSJ-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 8 hits for monomerid = 50055724
Found 8 hits for monomerid = 50055724
Affinity DataIC50: 82nMAssay Description:In vitro agonist binding affinity was determined by displacing the [3H]N-propylnorapomorphine ([3H]NPA) in rat striatal Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 88nMAssay Description:In vitro agonist binding affinity was determined by displacing the [3H]N-propylnorapomorphine ([3H]NPA) in rat striatal Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 412nMAssay Description:Displacement of [3H]spiperone from rat striatal Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 412nMAssay Description:Displacement of [3H]spiperone from rat striatal Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: 412nMAssay Description:In vitro binding affinity for dopamine receptor D2 on rat striatal membranes by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 704nMAssay Description:Displacement of [3H]spiperone from rat striatal Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:In vitro antagonist binding affinity was determined by displacing the [3H]-SCH- in rat striatal Dopamine receptor D1More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:In vitro antagonist binding affinity was determined by displacing the [3H]-SCH- in rat striatal Dopamine receptor D1More data for this Ligand-Target Pair