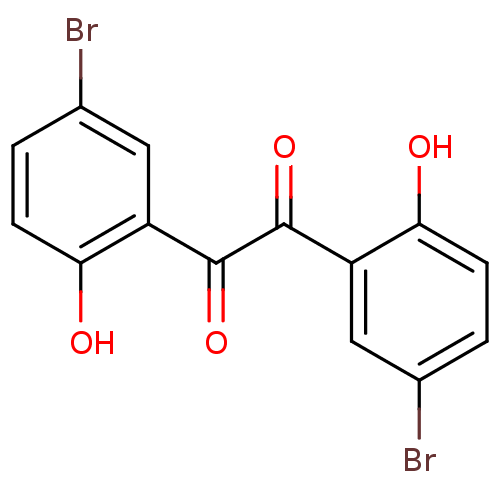
BDBM22753 1,2-bis(5-bromo-2-hydroxyphenyl)ethane-1,2-dione::Benzil-based compound, 31::cid_10662
SMILES Oc1ccc(Br)cc1C(=O)C(=O)c1cc(Br)ccc1O
InChI Key InChIKey=AAOYLOCWJSLLJU-UHFFFAOYSA-N
Data 5 KI
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 5 hits for monomerid = 22753
Found 5 hits for monomerid = 22753
Affinity DataKi: 53.5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: 72.7nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:CE inhibition was determined using a spectrophotometric multiwell plate assay with o-NPA as a substrate. The rate of change in absorbance at 420 nm w...More data for this Ligand-Target Pair