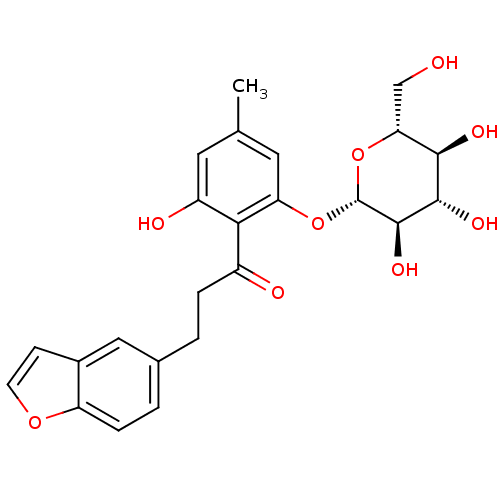
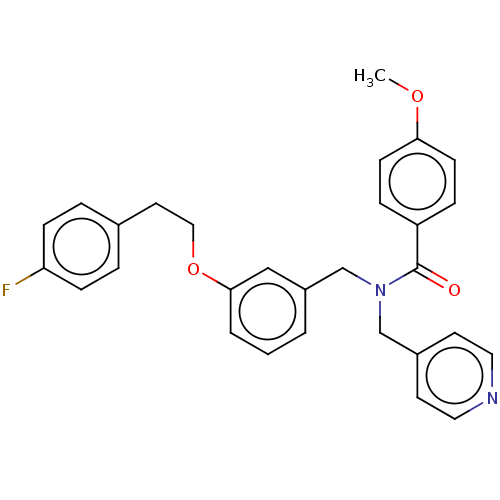
TargetSolute carrier family 2, facilitated glucose transporter member 1(Rat)
University of Kuopio
Curated by ChEMBL
University of Kuopio
Curated by ChEMBL
Affinity DataIC50: 710nMAssay Description:Inhibition of GluT1-mediated [14C]D-glucose uptake in Wistar rat brain by perfusion techniqueMore data for this Ligand-Target Pair
TargetSolute carrier family 2, facilitated glucose transporter member 1(Rat)
University of Kuopio
Curated by ChEMBL
University of Kuopio
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of GLUT1-mediated [3H]2-deoxy-glucose uptake in rat L6 cells after 15 mins by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSolute carrier family 2, facilitated glucose transporter member 1(Rat)
University of Kuopio
Curated by ChEMBL
University of Kuopio
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of GLUT1-mediated [3H]2-deoxy-glucose uptake in rat L6 cells after 15 mins by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSolute carrier family 2, facilitated glucose transporter member 1(Rat)
University of Kuopio
Curated by ChEMBL
University of Kuopio
Curated by ChEMBL
Affinity DataIC50: 3.29E+4nMAssay Description:Inhibition of GluT1-mediated [14C]D-glucose uptake in Wistar rat brain by perfusion techniqueMore data for this Ligand-Target Pair
TargetSolute carrier family 2, facilitated glucose transporter member 1(Rat)
University of Kuopio
Curated by ChEMBL
University of Kuopio
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Antagonist activity at rat C-terminal 6His/Myc-tagged GLUT1 assessed as reduction in 2-[3H]deoxyglucose uptake measured after 6 minsMore data for this Ligand-Target Pair